Genetic Association of GDF5, COG5 and CYBA Gene Polymorphisms with Primary Knee Osteoarthritis in South Indian Population
Hasan Q1* Poornima S1, Subramanyam K2, Khan IA3, Sumanlatha G4
1Department of Genetics and Molecular Medicine, Kamineni Hospitals, Hyderabad, Telangana, India
2Department of Orthopedics, Yashoda Hospitals, Hyderabad, Telangana, India
3Department of Clinical Laboratory Sciences, College of Applied Medical Sciences, King Saud University, Riyadh, Saudi Arabia
4Department of Genetics, Osmania University, Hyderabad, Telangana, India.
- *Corresponding Author:
- Dr. Qurratulain Hasan
Department of Genetics & Molecular Medicine
Kamineni Hospitals, LB Nagar
Hyderabad 500068, India
Tel: 040-39879999
E-mail: qhasan2000@yahoo.com
Received date: February 24, 2018; Accepted date: March 22, 2018; Published date: April 03, 2018
Copyright: © 2018 Poornima S, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Abstract
Purpose: Osteoarthritis (OA) is also known as degenerative arthritis and it affects millions of people and can have a huge impact on their lives. Single Nucleotide Polymorphisms (SNPs) in combination with genome wide association studies (GWAS) have been significantly focused. GWAS have been evaluated using candidate genes with genetic variants for Growth Differentiation Factor 5 (GDF5), Component of Oligomeric Golgi complex 5 (COG5), and Cytochrome B-245 alpha chain (CYBA) with primary knee Osteoarthritis (OA). These studies have been documented in Caucasian population, and we aimed to investigate the case-control study to evaluate association between knee OA in the South Indian population.
Materials & methods: SNPs for GDF5 (rs143383), COG5 (rs4730250) and CYBA (rs4676) were genotyped with polymerase chain reaction and restriction fragment length polymorphism in 150 primary Knee OA cases and 150 age and gender-matched controls. Alleles and genotype frequencies were calculated to compare the significant and non-significant association with the p-values. Multifactor dimensionality reduction (MDR) analysis was performed to study the gene-gene and gene-environment interactions.
Results: Baseline clinical details such as Body Mass Index (BMI), weight, histories of Type 2 diabetes and hypertension were significantly associated with primary Knee OA (p<0.05). Genetic factors for 3 SNPs were significantly associated with allelic frequency, dominant, and recessive mode of inheritance pattern (p<0.05). MDR analysis revealed a strong synergistic interaction between GDF5 and CYBA as well as COG5 and CYBA gene polymorphisms. GDF5 gene polymorphism is also synergistically interacting with BMI.
Conclusion: The three gene polymorphisms of GDF5, COG5 and CYBA could be used as a molecular biomarker to identify primary Knee OA in south Indian population to assess the risk of disease which in turn helps us to reduce the disability caused by the disease. To the best of our knowledge this is the first study in India.
Keywords
Osteoarthritis; GDF5; COG5; CYBA; South Indian population
Abbreviations
OA-Osteoarthritis; GWAS-Genome-Wide Association Studies ; GDF5-Growth Differentiation Factor 5; COG5-Component of an Oligomeric Complex; CYBA- Cytochrome B-245 Alpha Chain; SNPs-Single Nucleotide Polymorphisms; BMI-Body Mass Index; CDMP1-Cartilage-Derived Morphogenetic Protein 1; PCRPolymerase Chain Reaction; RFLP-Restriction Fragment Length Polymorphism; MDR-Multifactor Dimensionality Reduction; OROdds ratio; CI-Confidence Intervals; T2DM-Type 2 Diabetes Mellitus.
Introduction
Knee Osteoarthritis is commonly documented form for knee joint diseases in all the ages and considered mainly in the middle-age [1]. Osteoarthritis (OA) is one of the multifactorial diseases linked up with age and characterized through the loss of articular cartilage in synovial joints [2]. OA is the leading cause of chronic disability, in the elderly population and is characterized by joint pain, swelling, stiffness and is known for its association with poor quality of life [3,4]. The exact mechanisms responsible in the etiology of OA are still unclear, and multiple lifestyle and environmental factors are considered to significantly contribute to its risk. Age, weight, climate and economic status etc [5,6]. The increasing obesity is a growing burden in the elderly population because the knee joint is commonly affected by OA [2,7]. The results of heritability studies in twins, sibling pairs, and families have shown that genetic factors are estimated to contribute 50% of the risk for developing OA [8,9]. The role of specific genes is under investigation as the estimated heritability of primary OA is high, i.e., 40% for the knee, 60% for the hip, and 65% for the hand [10].
The investigations using Genome-Wide Association Studies (GWAS) have provided significant growth in the knowledge of genetics in OA [11,12], and multiple genes have been identified which are associated with complex diseases. Apart from GWAS, meta-analysis, linkage analysis, and candidate gene studies also implicated some genes in the etiology of OA [13]. In the present case-control study, three genes which play a role in the pathophysiology of knee OA were evaluated. Growth Differentiation Factor 5 (GDF5; ID-8200, MIM-601146), gene variant (rs143383), mapped to the gene cluster on chromosome 7 which includes the component of Oligomeric Golgi complex 5 (COG5; ID-10466, MIM-606821) gene and rs4676 (C242T) polymorphism is appeared as a functional variant in the promoter region of Cytochrome B-245, Alpha Polypeptide (CYBA; ID-1535; MIM-608508) have been identified as candidate genes for primary knee OA [14]. The GDF5 mutation has been associated with skeletal dysplasia’s, including Hunter-Thompsontype acromesomelic dysplasia, type C brachydactyly, and Grebetype chondrodysplasia in the humans and these associations endorse that GDF5 plays an important role in human skeletal development and maintenance [15]. GDF5 is also known as CDMP1 (cartilage-derived morphogenetic protein1), a growth factor with high articular cartilage and a member of BMP (bone morphogenetic protein) family of the Transforming Growth Factor β superfamily. The GDF5 is involved in the joint formation and expressed in the region of future joints during early development. The role of GDF5 in the pathogenesis of OA is known as it plays a key role in bone and cartilage morphogenesis, as well as, in joint formation [16]. The rs143383 polymorphism in the 5’-untranslated regions of GDF5 is associated with knee OA in Europeans and Asians [17,18]. The COG5 protein is one of eight proteins (Cog1-8) which form a Golgi-localized complex required for normal Golgi morphology and function. The encoded protein is organized with conserved oligomeric Golgi complex components 6,7 and eight into a subcomplex referred to as lobe B. Alternative splicing results in multiple transcript variants. Mutations in this gene result in congenital disorder of glycosylation type 2I [19]. CYBA gene encodes the light, alpha subunit which has been proposed as a primary component of the microbicidal oxidase system of phagocytes. CYBA/p22phox variants are associated with autosomal recessive chronic granulomatous disease, characterized by the failure of activated phagocytes to generate superoxide, which is important for the microbicidal activity of these cells [20]. Presently, there are no pharmacological interventions for OA patients. Replacement of knee joint is the only effective treatment at a severe stage of the osteoarthritis but is relatively costly and highly invasive. Based on the previous studies, we assume that multiple genes may be involved in susceptibility to knee OA and we designed a case-control study to evaluate the association between primary knee OA in South Indian population with three selected SNPs relevant in the pathogenesis of knee OA.
Materials and Methods
Sample selection
The present case-control study comprised 150 primary Knee OA cases which were clinically diagnosed, age and gendermatched 150 individuals with no musculoskeletal problems were considered as controls. Approval for the study was granted by the Institutional Ethics committee as per the 1964 Helsinki declaration and its later amendments or comparable ethical standards. After obtaining the signed informed consent from all the subjects, 2 ml of peripheral blood sample was drawn in EDTA vacutainer. Demographic details, along with clinical, personnel and family history were obtained from patient or patient’s care taker. In this hospital-based study, we have selected 150 primary knee OA cases from the Department of Orthopedics based on the inclusion and exclusion criteria described in our recent publication [21]. The primary knee OA cases were clinically diagnosed and radiologically confirmed with primary osteoarthritis as per the Kellegren/Lawrence score (0-4 score) [22].
Simultaneously, we have selected age, and gender-matched 150 controls from master health checkup based on questionnaire without any history of musculoskeletal diseases. The selection, inclusion and exclusion criteria of cases and controls have been described in detail in our prior publication [23].
Selection of SNPs
In the current study, three gene polymorphisms were selected based on the literature, i.e., GDF5 (rs143383), COG5 DUS4L (rs4730250) and CYBA/p22phox (rs4673) [14,20]. The details of the genes and SNPs have been documented in table 1.
| Gene | SNP/rsnumber | Position | Forward primers | Reverse primers | Enzymes | Band Sizes |
|---|---|---|---|---|---|---|
| GDF5 | C1104T/ rs143383 | 35438203 | CTCCTTCAAGCCCTCAGTCA | GTAGCAGCAGAAGGAAAGGC | BsiEI | C-160/44bp T-204bp |
| COG5 | A2850G/rs4730250 | 107567250 | AGAGTGACTGCATGCAAACG | ATGGTGGGTGGTCTTACAGT | DraI | A-174/26bp G-200bp |
| CYBA | C242T/rs4673 | 106994931 | TGCTTGTGGGTAAACCAAGGCCGGTG | AACACTGAGGTAAGTGGGGGTGGCTCCTGT | RsaI | C-188/160bp T-348bp |
Table 1: Information on selected variants involved in this study.
Genomic DNA was isolated from the peripheral blood sample using salting-out technique routinely used in our NABL accredited laboratory [24]. The quality of DNA was quantified using NanoDrop 2000 (Thermo Fisher Scientific, USA). The SNPs rs143383, rs4730250 and rs4673 were evaluated by polymerase chain reaction followed by restriction fragment length polymorphism and agarose gel electrophoresis using the specific primers described in table 1. The PCR products were digested with a BsiE1 restriction enzyme for rs143383 polymorphism, with Dra1 rs4730250 and with Rsa1 for rs4673. The digested products were run on 2.5% agarose gel staining with gel green dye to obtain the specific products after digestion. Genotype fragments were imaged using gel documentation system (Kamineni Life Sciences, Hyderabad, India).
Statistical analysis
Allele frequencies and genotype distribution of primary knee OA cases and controls were compared using Chi-square test. Med calc Version 12.7.0.0 was used to estimate odds ratio (OR) and 95% confidence intervals (CI) for an association of GDF5 (rs143383), COG5 (rs4730250) and CYBA (rs4673) genotype and the allele with primary knee OA. Multifactor dimensionality reduction (MDR) analysis was performed to study the gene-gene and gene-environment interactions.
Results
Baseline characteristics
This hospital-based Indian population study comprised of 300 samples. 150 primary knee OA cases (females-95 and males-55) between the ages of 28-50 years. The age and gender-matched 150 (females 90 and males 60) controls were included in the study. Clinical characteristics of 300 subjects at baseline are represented in table 2.
| Characters | Cases (n=150) | Controls (n=150) | p Value |
|---|---|---|---|
| Age (Years) | 45.09±8.83 | 46.46±7.77 | P=0.2538 |
| Sex (M:F) | 55:95 | 60:90 | P=0.5728 |
| Height (cm) | 155.78±3.19 | 155.48±4.18 | P=4.852 |
| Weight (kg) | 74.76±8.75 | 61.22±7.35 | P=0.0001 |
| BMI (kg/m2) | 30.81±3.49 | 25.38±3.37 | P<0.0001 |
| Age of Onset | 42.40±8.07 | NA | NA |
| Family History of OA | 32% | NA | NA |
| History of Hypertension | 57% | 16% | P=0.001 |
| History of T2DM | 36% | 11% | P=0.0056 |
| History of Thyroid Dysfunction | 28% | 13% | P=0.1254 |
Table 2: Clinical characteristics of primary knee OA cases and controls.
There is no significant difference between the mean height of the cases and controls (p=4.85). BMI (30.81±3.49) showed a significant difference between cases and control group (p<0.05). The history of T2DM and HTN were significantly associated with primary knee OA (p<0.05) and thyroid dysfunction was similar between cases and controls (p>0.05). A positive family history of primary knee OA was seen in 32% of cases.
Genetic factors
• Association of GDF5 gene polymorphism
The genotype distribution and allele frequencies of GDF5 (rs143383), COG5 (rs4730250) and CYBA (rs4673) variants were represented in table 3. Genotypic and allelic distributions of the GDF5 polymorphism follow the HWE in our controls. The variant T allele frequency in cases was 0.63 whereas in controls 0.37. The data indicated that the T allele and TT genotype was present in a higher percentage of cases compared to controls (Table 3). The heterozygous CT genotype was also comparatively high in cases (30%) whereas it is 23% controls. The data indicated a significant association of dominant (OR=3.73 95% CI=2.26–6.17, p<0.0001), and recessive mode of inheritance pattern (OR=2.62, 95% CI=1.61–4.26, p <0.0001). Furthermore, the variant allele T was significantly associated with the disease (OR=2.85, 95% CI=2.05–3.98, p<0.0001) which indicates that individuals with T allele have 2.85-fold increased risk for the development of primary knee OA in our population.
| rs number | Model | Genotypes | Cases | Controls | OR (95% CI) | p Value |
|---|---|---|---|---|---|---|
| rs143383 (GDF5) | Normal | CC | 33 (22%) | 77 (51.3%) | Reference | |
| Heterozygous | CT | 45 (30%) | 34 (22.7%) | 3.12 (1.84-5.26) | p<0.0001 | |
| Variant | TT | 72 (48%) | 39 (26%) | 1.31 (0.78-2.19) | p=0.29 | |
| Dominant | TT+CT vs. CC | 117(78%) | 73 (48.7%) | 3.73 (2.26-6.17) | p<0.0001 | |
| Co-dominant | CT vs. CC+TT | 45 (30%) | 34 (22.7%) | 1.33 (0.79-2.24) | p<0.27 | |
| Recessive | TT vs. CT+CC | 72 (48%) | 39 (26%) | 2.62 (1.61-4.26) | p<0.0001 | |
| C | 111(0.37) | 188(0.63) | Reference | |||
| Risk | T | 189 (0.63) | 112(0.37) | 2.85 (2.05-3.98) | p<0.0001 | |
| rs4730250 (COG5) | Normal | AA | 37 (24.6%) | 87 (58%) | Reference | |
| Heterozygous | AG | 44 (29.3%) | 33 (22%) | 3.23 (1.93-5.40) | p<0.0001 | |
| Variant | GG | 69 (46%) | 30 (20%) | 1.57 (0.91-2.69) | p=0.09 | |
| Dominant | GG+AG vs. AA | 113 (75.3%) | 63 (42%) | 4.21 (2.57-6.90) | p<0.0001 | |
| Co-dominant | AG vs. AA+GG | 44 (29.3%) | 33 (22%) | 1.47 (0.87-2.48) | p=0.14 | |
| Recessive | GG vs. AG+AA | 69 (46%) | 30 (20%) | 3.40 (2.04-5.69) | p<0.0001 | |
| A | 118 (0.39) | 207 (0.69) | Reference | |||
| Risk | G | 182 (0.61) | 93 (0.31) | 3.43 (2.45-4.80) | P<0.0001 | |
| rs4673 (CYBA) | Normal | CC | 35 (23.3%) | 105 (70%) | Reference | |
| Heterozygous | CT | 28 (18.7%) | 15 (10%) | 5.78 (3.06-10.91) | p<0.0001 | |
| Variant | TT | 87 (58%) | 30 (20%) | 1.52 (0.79-2.92) | p=0.20 | |
| Dominant | TT+CT vs. CC | 115 (76.7%) | 45 (30%) | 7.66 (4.58-12.82) | p<0.0001 | |
| Co-dominant | CT vs. CC+TT | 28 (18.7%) | 15 (10%) | 2.06 (1.05-4.04) | p=0.03 | |
| Recessive | TT vs. CT+CC | 87 (58%) | 30 (20%) | 5.52 (3.30-9.24) | p<0.0001 | |
| C | 98 (0.32) | 225 (0.75) | Reference | |||
| Risk | T | 202 (0.68) | 75 (0.25) | 6.18 (4.33-8.82) | p<0.0001 |
Table 3: Statistical association of SNPs with OA cases disease risk compared with controls.
• Association of COG5 gene polymorphism
Genotypic and allelic distribution of COG5 gene polymorphism follows the HWE in our population. The variant G allele of the SNP rs4730250 in COG5 gene had the frequency 0.61 in cases whereas it was 0.31 in controls. The genotype frequencies of AA, AG, and GG are documented as 25%, 29%, 46% in primary Knee OA cases and 58%, 22%, 20% in controls respectively. The data indicated a significant association of dominant (OR=4.21, 95% CI=2.57–6.93, p<0.001) and recessive (OR=3.40, 95% CI=2.04–5.69, p<0.0001) modes of inheritance with the disease pathology. Furthermore, the variant allele G also was significantly associated with primary knee OA (OR=3.43, 95% CI=2.45–4.80, p<0.0001) (Table 3).
• Association of CYBA gene polymorphism
Genotypic and allelic distributions of the CYBA polymorphism satisfied the HWE in our study group. Genotypic and allelic frequencies are tabulated in table 3. The variant T allele in cases was 0.68 and in controls was 0.25. The data indicated that the T allele was present in a higher percentage of cases (58%) compared to controls (20%). The data indicated a significant association of dominant (OR=7.66, 95% CI=4.58–12.82, p<0.0001), co-dominant (OR=2.06, 95% CI=1.05–4.04, p=0.034), and recessive (OR=5.52, 95% CI=3.30 – 9.24, p<0.0001) modes of inheritance with primary knee OA. Furthermore, the variant allele T was statistically significant with the disease pathology (OR = 6.18, 95% CI=4.33–8.82, p<0.0001) (Table 3).
Genotypes association between males and females
There is no significant difference between males and females of the three candidate gene polymorphisms analyzed in this study with primary knee osteoarthritis (Table 4).
| rs number | Model | Genotypes | Male | Female | OR (95% CI) | p Value |
|---|---|---|---|---|---|---|
| rs143383 (GDF5) | Normal | CC | 11 (20%) | 22 (23%) | Reference | |
| Heterozygous | CT | 19 (35%) | 26 (27%) | 1.38 (0.62-3.07) | p=0.42 | |
| Variant | TT | 25 (45%) | 47 (50%) | 0.91 (0.47-1.75) | p=0.77 | |
| Dominant | TT+CT vs. CC | 44 (80%) | 73 (77%) | 1.20 (0.53-2.7) | p=0.65 | |
| Co-dominant | CT vs. CC+TT | 19 (35%) | 26 (27%) | 1.40 (0.68-2.86) | p=0.35 | |
| Recessive | TT vs. CT+CC | 25 (45%) | 47 (50%) | 0.85 (0.43-1.65) | p=0.63 | |
| C | 67 (0.37) | 70 (0.37) | Reference | |||
| Risk | T | 69 (0.63) | 120 (0.63) | 0.60 (0.38-0.93) | p=0.02 | |
| rs4730250 (COG5) | Normal | AA | 19 (35%) | 18 (19%) | Reference | |
| Heterozygous | AG | 13 (24%) | 31 (33%) | 0.53 (0.25-1.15) | p=0.11 | |
| Variant | GG | 23 (41%) | 46 (48%) | 1.18 (0.59-2.33) | p=0.62 | |
| Dominant | GG+AG vs. AA | 36 (65%) | 77 (81%) | 0.44 (0.20-0.94) | p=0.03 | |
| Co-dominant | AG vs. AA+GG | 13 (24%) | 31 (33%) | 0.63 (0.30-1.36) | p=0.24 | |
| Recessive | GG vs. AG+AA | 23 (41%) | 46 (48%) | 0.76 (0.39-1.49) | p=0.43 | |
| A | 51 (0.46) | 67 (0.35) | Reference | |||
| Risk | G | 59 (0.54) | 123 (0.65) | 0.63 (0.39-1.01) | p=0.05 | |
| rs4673 (CYBA) | Normal | CC | 11 (20%) | 24 (25%) | Reference | |
| Heterozygous | CT | 9 (16%) | 19 (20%) | 1.22 (0.50-2.97) | p=0.65 | |
| Variant | TT | 35 (64%) | 52 (55%) | 1.16 (0.55-2.45) | p=0.69 | |
| Dominant | TT+CT vs. CC | 44 (80%) | 71 (75%) | 1.35 (0.60-3.02) | p=0.46 | |
| Co-dominant | CT vs. CC+TT | 9 (16%) | 19 (20%) | 0.78 (0.32-1.87) | p=0.58 | |
| Recessive | TT vs. CT+CC | 35 (64%) | 52 (55%) | 1.44 (0.73-2.86) | p=0.28 | |
| C | 31 (0.28) | 67 (0.35) | Reference | |||
| Risk | T | 79 (0.72) | 123 (0.65) | 1.38 (0.83-2.31) | p=0.20 |
Table 4: Statistical association between men and female genotypes in OA cases.
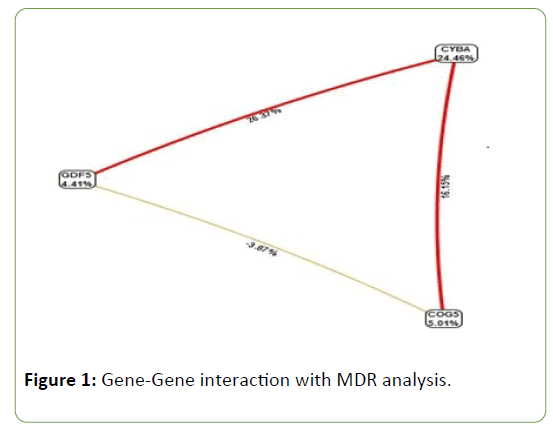
Gene-Gene interactions
MDR is a powerful statistical tool for detecting and modeling epistasis. The interaction graph indicates that GDF5, COG5, and CYBA polymorphisms contribute 4.41%, 5.01%, and 24.46%, respectively, towards the pathology of primary knee OA (Figure 1).
GDF5 and CYBA (23.37%) as well as COG5 and CYBA (16.15%) polymorphisms were found to have a strong synergistic interaction whereas GDF5 and COG5 (-3.87%) polymorphism showed negative interaction.
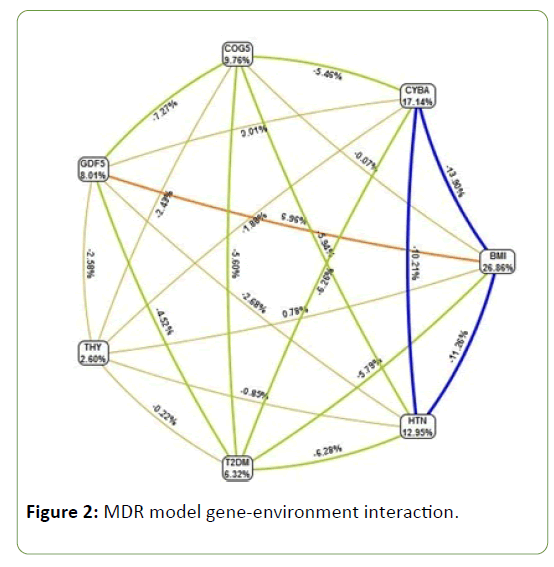
Gene–Environment interactions
MDR analysis was also performed to find interactions among genotypes between cases and controls, as well as among demographic parameters. Among demographic parameters, BMI (26.86%) contributed highest to disease pathology. GDF5 gene polymorphism and BMI showed strong synergistic interaction (56.9%). Comorbid factors like hypertension contributed 12.95%, diabetes 6.32% and thyroid dysfunction 2.6 % (Figure 2).
Figure 2: MDR model gene-environment interaction.
Discussion
Osteoarthritis, is a phenomenon occurring in previously intact joints with no apparent initiating factor such as joint injury or developmental abnormalities. Gene identification is important for understanding the disease pathophysiology for improving diagnosis, prevention, and treatment. The underlying genetic background could provide new insights into the pathophysiology of OA which could potentially lead to new therapeutic targets [10,24]. The core purpose of this study was to investigate the association between GDF5 (rs143383), COG5 (rs4730250) and CYBA (rs4676) polymorphisms as potential genetic biomarkers for identifying individuals who are at risk of developing primary knee OA in a cosmopolitan city of the south India. We found that T allele of GDF5 and CYBA gene polymorphisms and G allele of COG5 gene showed a strong significant association with primary knee OA cases. To the best of our knowledge, this is the first association study carried out with three SNPs in south Indian population.
GDF5 gene regulates the expression of a GDF5 protein. It involves in the development of bone and cartilage, particularly in endochondral classification [25]. Recently several studies have shown an association between Growth Differentiation Factor 5 polymorphism and knee osteoarthritis, but the crucial role of predisposing genetic factor in each ethnic group cannot be replicated in all populations. The first association of GDF5 and OA was reported in Japanese [18] and reproduced in Chinese and European Caucasians [15]. In India, this is the first study which reported an association of GDF5 gene polymorphism with knee osteoarthritis. The rs143383 polymorphism was evaluated in different ethnic population, and several meta-analyses have been carried out and concluding it to be significant [6,26,27]. The association of this polymorphism from Korean and Greek population showed negative results [28,29]. GDF5 rs143383 gene polymorphism and knee osteoarthritis have suggested stronger association in Asians than Caucasians [13].
Tawonsawtuk et al. study revealed that GDF5 gene polymorphism has an association with knee OA in Thai ethnic population. A Northern European study demonstrated that an alliance of Lumbar Disc degeneration with T allele of GDF5 gene polymorphism as in knee OA and hip OA [30]. The ethnicity based studies appears to be positive association [12,15,17,31-35] and negative association [29,36-39]. However, a couple of studies have been carried out in female subjects and turn to be similar [16] and dissimilar association [40], this study was carried out the rs4730250 polymorphism in Finnish female OA subjects and confirms the significant association.
The gene COG5 was identified from an arcOGEN study. The association of this locus with knee OA was discovered in a GWAS which after different study phases of 500,000 SNPs led to a single SNPs, rs3815148, with a GWAS level of association (OR=1.14, p=8×10−8). To the best of our current knowledge, this is the first study in India that showed a significant association between primary knee OA with G allele of COG5 gene. The weak correlation found with patellofemoral radiographic grade, suggest that this gene might play a role in joint damage and indicates the need for studies on functional relevance [14]. Raine et al. [41] fail to confirm the statistical association and Evangelou et al. [42] carried out a meta-analysis of GWAS studies confirms susceptibility locus for knee OA on the 7q22 chromosome.
In CYBA gene, the C242T polymorphism results in a substitution of Tyr for His at residue 72 of p22phox, and significantly r educed v ascular NADPH o xidase activity [43]. It might be expected to reduce the generation of ROS, indicating a protective role of T allele in ACS, which is consistent with the results for the Asian population as indicated in the meta-analysis [44]. We can identify a single study carried out with the rs4676 polymorphism in Greek population and confirms negative association with OA [20].
High BMI was established as a risk factor for OA patients. We have categorized OA cases based on BMI into (i) normal (18.5– 24.9 BMI), (ii) overweight (25–29.9 BMI) and (iii) obese (>30 BMI) groups. In GDF5 gene polymorphism the percentage of TT genotype in obese and overweight cases were 48% whereas CT genotype was 32% and 26% in obese and overweight respectively which is higher when compared with CC genotype of obese (20%) and overweight (26%) population. TT genotype was playing a role in disease pathology. In COG5 gene polymorphism the variant GG genotype and heterozygous AG genotype percentage was higher in over weight and obese population when compared with the people who had normal BMI. GG genotype was playing a role in disease pathology, and obese individuals with T allele are more prone for primary knee OA. In CYBA gene polymorphism the percentage of TT genotype was 64% in obese which is higher than the individuals with normal (60%) BMI. Obese individuals with TT genotype were more prone to the disease and individuals with TT genotype irrespective of BMI are at increased risk for the disease (Table 5). The strength of this study was in selecting the right samples of both primary knee OA cases and controls in the hospital set up and perform the PCR-RFLP SNPs in three different genes which were associated with disease pathology. The current limitation of our study may be small sample size with single SNP from each candidate gene.
| BMI ranges | OA Cases (n=150) |
BMI (Mean±SD) |
Genotypes-GDF5 | ||
|---|---|---|---|---|---|
| CC | CT | TT | |||
| Normal (0-24.9) | 10 | 23.12±1.10 | 2 (20%) | 3(30%) | 5(50%) |
| Overweight (25.0-29.9) | 46 | 27.96±1.30 | 12 (26%) | 12 (26%) | 22 (48%) |
| Obese (>30.0) | 94 | 33.02±1.83 | 19(20%) | 30(32%) | 45(48%) |
| OA Cases (n=150) |
BMI (Mean±SD) |
Genotypes-COG5 | |||
| AA | AG | GG | |||
| Normal (0-24.9) | 10 | 23.12±1.10 | 3 (30%) | 3(30%) | 4(40%) |
| Overweight (25.0-29.9) | 46 | 27.96±1.30 | 10(22%) | 12 (26%) | 24 (52%) |
| Obese (>30.0) | 94 | 33.02±1.83 | 24(26%) | 29(31%) | 41(43%) |
| OA Cases (n=150) |
BMI (Mean±SD) |
Genotypes-CYBA | |||
| CC | CT | TT | |||
| Normal (0-24.9) | 10 | 23.12±1.10 | 1 (10%) | 3(30%) | 6(60%) |
| Overweight (25.0-29.9) | 46 | 27.96±1.30 | 16 (35%) | 09 (20%) | 21 (45%) |
| Obese (>30.0) | 94 | 33.02±1.83 | 18(19%) | 16(17%) | 60(64%) |
Table 5: Variance with BMI and genotypes of GDF5, COG5 & CYBA genes in this study.
Conclusion
In conclusion, this initial study explores the genetic association of GDF5, COG5 and CYBA gene polymorphisms with primary knee OA in the South Indian population and could be used as molecular biomarkers to identify the individuals who are at risk of developing primary knee OA. To the best of our knowledge, this first study in South Indian population explored significant association.
Author’s contribution
PS has carried out the research experiment and written the manuscript; SK is a clinician and has helped us with the OA samples; KIA edited the manuscript for the better quality. GS is a Co-I of the project and analysed the data; and HQ is the PI of the project, designed the study, analyzed the data and edited and finalized the manuscript.
Conflict of Interest
There is no conflict of interest towards this article.
Acknowledgement
The authors extend their sincere appreciation to Kamineni Hospitals, Hyderabad towards financial support and also for providing equipment and infrastructure to carry out the experimental work. We also extend our gratitude towards the subjects involved in this study.
Funding source
We did not receive any funding from any of the funding agencies.
References
- Liu R, Yuan X, Yu J, Quan Q, Meng H, et al. (2017) An updated meta-analysis of the asporin gene D-repeat in knee osteoarthritis: effects of gender and ethnicity. J Orthop Surg Res 12: 148.
- Shepherd C, Skelton AJ, Rushton MD, Reynard LN, Loughlin J (2015) Expression analysis of the osteoarthritis genetic susceptibility locus mapping to an intron of the MCF2L gene and marked by the polymorphism rs11842874. BMC Med Genet 16: 108.
- Collins NJ, Prinsen CA, Christensen R, Bartels EM, Terwee CB, et al. (2016) Knee injury and Osteoarthritis Outcome Score (KOOS): Systematic review and meta-analysis of measurement properties. Osteoarthritis Cartilage 24: 1317-1329.
- Heidari B (2011) Knee osteoarthritis diagnosis, treatment and associated factors of progression: Part II. Caspian J Intern Med 2: 249-255.
- Wang Q, Yan XB, Sun QQ, Hu AM, Liu HL, et al. (2015) Genetic polymorphism of the estrogen receptor alpha gene and susceptibility to osteoarthritis: evidence based on 15,022 subjects. Curr Med Res Opin 31: 1047-1055.
- Zhang Q, Li H, Zhang Z, Yang F, Chen J (2015) Serum metabolites as potential biomarkers for diagnosis of knee osteoarthritis. Disease markers 2015: 684794.
- Salih S, Sutton P (2013) Obesity, knee osteoarthritis and knee arthroplasty: A review. BMC Sports Sci Med Rehabil 5: 25.
- Miyamoto Y, Shi D, Nakajima M, Ozaki K, Sudo A, et al. (2008) Common variants in DVWA on chromosome 3p24.3 are associated with susceptibility to knee osteoarthritis. Nat Genet 40: 994-998.
- Zhang R, Yao J, Xu P, Ji B, Luck JV, et al. (2015) A comprehensive meta-analysis of association between genetic variants of GDF5 and osteoarthritis of the knee, hip and hand. Inflamm Res 64: 405-414.
- Kerkhof HJ, Lories RJ, Meulenbelt I, Jonsdottir I, Valdes AM, et al. (2010) A genome-wide association study identifies an osteoarthritis susceptibility locus on chromosome 7q22. Arthritis Rheum 62: 499-510.
- Bravata V, Minafra L, Forte GI, Cammarata FP, Saporito M, et al. (2015) DVWA gene polymorphisms and osteoarthritis. BMC Res Notes 8: 30.
- Minafra L, Bravata V, Saporito M, Cammarata FP, Forte GI, et al. (2014) Genetic, clinical and radiographic signs in knee osteoarthritis susceptibility. Arthritis Res Therapy 16: R91.
- Pan F, Tian J, Winzenberg T, Ding C, Jones G (2014) Association between GDF5 rs143383 polymorphism and knee osteoarthritis: an updated meta-analysis based on 23,995 subjects. BMC Musculoskelet Disord 15: 404.
- Valdes AM, Doherty S, Muir KR, Zhang W, Maciewicz RA, et al. (2012) Genetic contribution to radiographic severity in osteoarthritis of the knee. Ann Rheum Dis 71: 1537-1540.
- Ikegawa S (2013) The genetics of common degenerative skeletal disorders: Osteoarthritis and degenerative disc disease. Annu Rev Genomics Hum Genet 14: 245-256.
- Xiao JL, Meng JH, Gan YH, Zhou CY, Ma XC (2015) Association of GDF5, SMAD3 and RUNX2 polymorphisms with temporomandibular joint osteoarthritis in female Han Chinese. J Oral Rehabil 42: 529-536.
- Chapman K, Takahashi A, Meulenbelt I, Watson C, Rodriguez-Lopez J, et al. (2008) A meta-analysis of European and Asian cohorts reveals a global role of a functional SNP in the 5′ UTR of GDF5 with osteoarthritis susceptibility. Hum Mol Genet 17: 1497-1504.
- Miyamoto Y, Mabuchi A, Shi D, Kubo T, Takatori Y, et al. (2007) A functional polymorphism in the 5' UTR of GDF5 is associated with susceptibility to osteoarthritis. Nat Genet 39: 529-533.
- Rodriguez-Fontenla C, Gonzalez A (2015) Genetics of osteoarthritis. Reumatologia Clinica 11: 33-40.
- Lepetsos P, Pampanos A, Lallos S, Kanavakis E, Korres D, et al. (2013) Association of NADPH oxidase p22phox gene C242T, A640G and -930A/G polymorphisms with primary knee osteoarthritis in the Greek population. Mol Biol Rep 40: 5491-5499.
- Subramanyam K, Poornima S, Juturu KK, Anand D, Mohanthy S, et al. (2016) Missense FokI variant in the vitamin D receptor gene in primary knee osteoarthritis patients in south Indian population. Gene Reports 4: 118-122.
- Kellgren JH, Lawrence JS (1957) Radiological assessment of osteo-arthrosis. Ann Rheumatic Dis 16: 494-502.
- Poornima S, Subramanyam K, Khan IA, Hasan Q (2015) The insertion and deletion (I28005D) polymorphism of the angiotensin I converting enzyme gene is a risk factor for osteoarthritis in an Asian Indian population. J Renin Angiotensin Aldosterone Syst 16: 1281-1287.
- Khan IA, Vattam KK, Jahan P, Hasan Q, Rao P (2016) Importance of glucokinase -258G/A polymorphism in Asian Indians with post-transplant and type 2 diabetes mellitus. Intract Rare Dis Res 5: 25-30.
- Hatakeyama Y, Tuan RS, Shum L (2004) Distinct functions of BMP4 and GDF5 in the regulation of chondrogenesis. J Cellular Biochem 91: 1204-1217.
- Hao SW, Jin QH (2013) Association between the +104T/C polymorphism in the 5'UTR of GDF5 and susceptibility to knee osteoarthritis: a meta-analysis. Molecular Med Reports 7: 485-488.
- Liu J, Cai W, Zhang H, He C, Deng L (2013) Rs143383 in the growth differentiation factor 5 (GDF5) gene significantly associated with osteoarthritis (OA)-a comprehensive meta-analysis. Int J Med Sci 10: 312-319.
- Cao Z, Lee HS, Song JH, Yoon JW, Park YK, et al. (2010) Growth differentiation factor 5 (GDF5) core promoter polymorphism is not associated with susceptibility to osteoarthritis of the knee in the Korean population. Korean J Pathol 44: 448-452.
- Tsezou A, Satra M, Oikonomou P, Bargiotas K, Malizos KN (2008) The growth differentiation factor 5 (GDF5) core promoter polymorphism is not associated with knee osteoarthritis in the Greek population. J Ortho Res 26: 136-140.
- Williams F, Popham M, Hart D, de Schepper E, Biermaâ€ÂÂZeinstra S, et al. (2011) GDF5 singleâ€ÂÂnucleotide polymorphism rs143383 is associated with lumbar disc degeneration in Northern European women. Arthritis Rheum 63: 708-712.
- Ikegawa S (2008) Genomic approaches to bone and joint diseases. Current status of genetic study of osteoarthritis. Clin Calcium 18: 162-167.
- Reynard LN, Bui C, Syddall CM, Loughlin J (2014) CpG methylation regulates allelic expression of GDF5 by modulating binding of SP1 and SP3 repressor proteins to the osteoarthritis susceptibility SNP rs143383. Hum Genet 133: 1059-1073.
- Tawonsawatruk T, Changthong T, Pingsuthiwong S, Trachoo O, Sura T, et al. (2011) A genetic association study between growth differentiation factor 5 (GDF 5) polymorphism and knee osteoarthritis in Thai population. J Ortho Surg Res 6: 47.
- Vaes RB, Rivadeneira F, Kerkhof JM, Hofman A, Pols HA, et al. (2009) Genetic variation in the GDF5 region is associated with osteoarthritis, height, hip axis length and fracture risk: the Rotterdam study. Ann Rheum Dis 68: 1754-1760.
- Valdes AM, Spector TD, Doherty S, Wheeler M, Hart DJ, et al. (2009) Association of the DVWA and GDF5 polymorphisms with osteoarthritis in UK populations. Ann Rheum Dis 68: 1916-1920.
- Bijsterbosch J, Kloppenburg M, Reijnierse M, Rosendaal FR, Huizinga TW, et al. (2013) Association study of candidate genes for the progression of hand osteoarthritis. Osteoarthritis Cartilage 21: 565-569.
- Dodd AW, Rodriguez-Fontenla C, Calaza M, Carr A, Gomez-Reino J, et al. (2011) Deep sequencing of GDF5 reveals the absence of rare variants at this important osteoarthritis susceptibility locus. Osteoarthritis Cartilage 19: 430-434.
- Egli RJ, Southam L, Wilkins JM, Lorenzen I, Pombo-Suarez M, et al. (2009) Functional analysis of the osteoarthritis susceptibility-associated GDF5 regulatory polymorphism. Arthritis Rheum 60: 2055-2064.
- Shin MH, Lee SJ, Kee SJ, Song SK, Kweon SS, et al. (2012) Genetic association analysis of GDF5 and ADAM12 for knee osteoarthritis. Joint Bone Spine 79: 488-491.
- Hamalainen S, Solovieva S, Vehmas T, Luoma K, Leino-Arjas P, et al. (2014) Genetic influences on hand osteoarthritis in Finnish women--a replication study of candidate genes. PLoS One 9: e97417.
- Raine EV, Wreglesworth N, Dodd AW, Reynard LN, Loughlin J (2012) Gene expression analysis reveals HBP1 as a key target for the osteoarthritis susceptibility locus that maps to chromosome 7q22. Ann Rheum Dis 71: 2020-2027.
- Evangelou E, Valdes AM, Kerkhof HJ, Styrkarsdottir U, Zhu Y, et al. (2011) Meta-analysis of genome-wide association studies confirms a susceptibility locus for knee osteoarthritis on chromosome 7q22. Ann Rheum Dis 70: 349-355.
- Guzik TJ, West NE, Black E, McDonald D, Ratnatunga C, et al. (2000) Functional effect of the C242T polymorphism in the NAD (P) H oxidase p22phox gene on vascular superoxide production in atherosclerosis. Circulation 102: 1744-1747.
- Hu P, Huang MY, Hu XY, Xie XJ, Xiang MX, et al. (2015) Meta-analysis of C242T polymorphism in CYBA genes: risk of acute coronary syndrome is lower in Asians but not in Caucasians. J Zhejiang Univ Sci 16: 370-379.
Open Access Journals
- Aquaculture & Veterinary Science
- Chemistry & Chemical Sciences
- Clinical Sciences
- Engineering
- General Science
- Genetics & Molecular Biology
- Health Care & Nursing
- Immunology & Microbiology
- Materials Science
- Mathematics & Physics
- Medical Sciences
- Neurology & Psychiatry
- Oncology & Cancer Science
- Pharmaceutical Sciences