Genetic Relatedness of Trichomonas vaginalis Isolates to the Clinical Variability
Wafaa A Abou-kamar1, Aida A Abdel-Mageid1, Hala A El-Nahas1, Raefa A ATIA1, Salwa A Abou Elkhair2 and Alaa A Mosbah3 , Nora L El-Tantawy1*
1Department of Medical Parasitology, Mansoura University, Mansoura, Egypt
2Department of Medical Biochemistry, Mansoura University, Mansoura, Egypt
3Department of Gynecology and Obstetrics, Mansoura University, Mansoura, Egypt
- *Corresponding Author:
- Nora L El-Tantawy
Assistant Professor
Department of Parasitology
Faculty of Medicine, Mansoura University, Egypt
Tel: +00201099931619
Fax: +20-50-226-3717
E-mail: noratantawy@yahoo.com; noralabeeb@mans.edu.eg
Received Date: 29 September 2017; Accepted Date: 19 October 2017; Published Date: 26 October 2017
Citation: Abou-kamar WA, Abdel-Mageid AA, El-Nahas HA, Atia RA, El-Tantawy NL, et al. (2017) Genetic Relatedness of Trichomonas vaginalis Isolates to the Clinical Variability. J Mol Microbiol. Vol. 1 No. 1: 103.
Copyright: © 2017 El-Tantawy NL, et al. This is an open-access article distributed under the terms of the creative Commons attribution License, which permits unrestricted use, distribution and reproduction in any medium, provided the original author and source are credited.
Abstract
Background: Trichomoniasis manifestation shows an extensive clinical variability ranging from no symptoms to severe presentation and sequel. An association is suggestive between the high genetic diversity of Trichomonas vaginalis isolates and its clinical variability. The aim of our study was to identify T. vaginalis isolates in symptomatic and asymptomatic females besides, studying their possible relation to different genotypes in our locality. Methods and findings: Two hundred female were included in the study and screened for T. vaginalis infection using the wet mount, culture in Diamond’s media and PCR amplification which targeted actin gene. For genotyping by RFLP, products of PCR amplification of actin gene were digested by 3 restriction enzymes (HindII, MseI and Rsal). Trichomonas vaginalis was detected in 12% (12/100) and 1% (1/100) in symptomatic and asymptomatic females respectively. But, by using modified Diamond culture and PCR, 22% (22/100) symptomatic and 5% (5/100) asymptomatic females were positive for T. vaginalis. After digestion of PCR products using restriction enzymes, different electrophoretic band patterns were demonstrated. Interpretation of these patterns, only one distinct electrophoretic pattern of actin genotype H in all studied isolates was demonstrated. Conclusion: Our results concluded that genetic variation does not correlate with the clinical variability of T. vaginalis as one genotype band pattern of actin gene was detected in both symptomatic and asymptomatic groups. Further studies in a large scale of isolates are needed to elucidate this finding.
Keywords
Trichomonas vaginalis; Manifestation; Genotyping; PCR; RFLP
Introduction
Trichomonas vaginalis is the most common preventable sexually transmitted infection and considered as one of the neglected tropical parasitic infections [1]. Trichomoniasis is presented with various clinical manifestations. More than 80% of patients are asymptomatic [2]. Common manifestations of acute infection include: vaginitis, vulval irritation and inflammation, malodorous vaginal discharge and punctate microhemorrhages on the cervix known as ‘strawberry cervix’ [3]. Infection with T. vaginalis frequently leads to inflammatory pelvic disease and infertility [4]. Moreover, infection with T. vaginalis predisposes to prostate cancer and acquisition of other infections like the human immunodeficiency virus (HIV) and Human papillomavirus (HPV) [5-7].
There is evidence regarding the correlation between T. vaginalis genotypes and clinical manifestations of trichomoniasis but, it is still not definite. Studies suggesting this correlation based on the matching between the extensive clinical variability in trichomoniasis and its sequel, and extensive clinical variability in trichomoniasis [5,8-10], Studies have shown concordance between T. vaginalis genotype and the clinical presentation of trichomoniasis, like susceptibility to metronidazole, presence of T. vaginalis virus, or concomitant Mycoplasma infection [11,12]. On the other hand, other studies based on RAPD and RFLP did not demonstrate any concordance between T. vaginalis genotypes and its symptoms [11,13].
Trichomonas vaginalis has a large genome of 160 Mb with approximately 60000 protein-coding genes. Most of its genome has repeats and transposable elements [14]. Actin protein of T. vaginalis is the main component of its cytoskeleton and has an essential role in cellular mobility and cell interaction [15]. This structural protein has a well- conserved ubiquitous nature, making it a feasible choice for intra-species molecular identification [16].
Commonly, PCR and its related methods were widely used for genetic studies in organisms. Random amplified polymorphic DNA (RAPD) method was used to reveal an association between T. vaginalis genotype and metronidazole resistance [8]. Also, restriction fragment length polymorphism (RFLP) and PCR-RFLP methods were applied to demonstrate the genetic diversity in clinical isolates of T. vaginalis [15,17]. PCR-RFLP has advantages of both the sensitivity of PCR and reliability of RFLP and so, it is considered as the best method for isolates genotyping [18,19].
The aim of this study was to identify T. vaginalis infection in both symptomatic and asymptomatic women and to study the possible correlation between T. vaginalis genotypes and clinical variability based on PCR-RFLP of actin gene.
Materials and Methods
Ethics statements
This study was approved by the Institutional Review Board of the faculty of medicine, Mansoura University, Egypt. Prior to enrollment, a signed consent was gained from each participant. Females with confirmed trichomoniasis were all treated and followed up.
Study participants
This study was carried out in Gynecology out-patients clinic of Mansoura University Hospital, Biochemistry and Parasitology department, faculty of medicine, Mansoura University, Egypt. Vaginal swabs were collected from 100 female with symptoms suggestive of vaginitis and another 100 asymptomatic female. The study was conducted in the period from December 2013 to February 2016. The age of the study participants ranged from 18-48 years. Females who were pregnant, received anti-parasitic or anti-trichomonal therapy, used vaginal wash within the previous two weeks were excluded from the study, Sociodemographic and clinical data were recorded.
Specimen collection, processing and culture
Enrolled females were examined in accordance with a standard clinical protocol. Three speculum-assisted vaginal swabs samples were collected from the posterior fornix of the vagina of each participant. The first swab was kept in a tube containing 3 ml sterile phosphate Buffered Saline (PBS) for wet mounts microscopy which is done immediately according to McCann JS [20]. Other two were transferred to a 10 ml prewormed screw-cap tube with sterile modified Diamond’s medium for culture. The medium was supplemented with streptomycin-penicillin at 50 μg/mL and 10% heat-inactivated bovine serum. Cultures were incubated at 37°C and examined for the presence of T. vaginalis by microscopy every day for 5 to 7 days post-inoculation.
DNA extraction and PCR amplification
Positive culture media was centrifuged and the pellet was washed twice in PBS at 4000 rpm for 10 min. and stored at -20°C. Genomic DNA was extracted from pellets by Thermos Scientific Gene JET Genomic DNA Purification Kit (USA) according to the manufacturer’s instructions. Quantity and purity of the extracted DNA was measured with Nanodrop (Thermo scientific NanoDrop 2000 spectrophotometer, USA) while, the quality of extracted DNA was tested by 1.5% agarose gel electrophoresis. The actin gene fragments were amplified.
Actin gene amplification: Nested PCR was employed and two sets of primers were used to amplify the (1100 bp) of T. vaginalis actin gene [21]. The outer and inner primers were selected from Crucitti T et al. [15]. The outer set primers were Tv8S Forward: 5’-TCTGGAATGGCTGAAGAAGACG-3’ and Tv9 Reverse: 5’-CAGGGTACATCGTATTGGTC-3’, the inner primers were Tv10S Forward: 5’-CAGACACTCGTTATCG-3’ and Tv11 Reverse: 5’-CGGTGAACGATGGATG-3’.
Reaction with outer primers: Each PCR reaction mixture contain 40 μl (20 μl of master mix, 10 μl of DNA template, 2 μl of forward outer primers, 2 μl of reverse outer primers and 6 μl sterile distilled water). PCR amplification was performed in two stages using thermocycler (TECHEN TC-312, FTC3102D, Barloworld Scientific Ltd. Stone, Stafford Shire, and UK). The first stage consist of 10 cycle, the first cycle was preceded by 5 min of denaturation at 95°C , then each cycle consisted of denaturation at 94°C for 30 sec, of annealing at 55°C for 30 sec, and extension at 72°C for 3 min. The second stage consisted of 25 cycles with the same denaturation and annealing steps. The extension step was extended by 5 seconds per cycle. The last cycle was followed by a 7 min final extension.
Reaction with inner primers: Using the same amplification program as outer primers with total reaction of 50 μl (25 μl master mix, 2 μl forward inner primers, 2 μl reverse inner primers, 4 μl PCR product and 17 μl DW).
The expected PCR products 1100 bp of amplified actin gene were confirmed by electrophoresis 2% (w/v) agarose gel, stained by ethidium bromide and visualized under an UV transilluminator.
Restriction fragment length polymorphism (RFLP)
For RFLP technique, amplicons of actin gene were digested by HindII, MseI and Rsal restriction enzymes. In brief, in each PCR tube, 17 μL of sterile distilled water, 2 μL (10X) of Fast digest buffer, 10 μL of amplified PCR product and 1μl of the restriction endonuclease are added. The reaction mixture was incubated at 37°C for 15 minutes then the enzyme was inactivated by heating for 5 min at 65°C in heat block. Finally, for visualization of product after digestion, the reaction was run on 2% agarose gel, run with 50 bp DNA markers at 100 volts for 45 minutes and then visualized via light UV transilluminator.
Statistical analysis
The data was plotted and analyzed using the statistical software package SPSS version 22.
Results
The total number of females involved in this study was 200, of whom 100 symptomatic and 100 were asymptomatic. The mean age was (35.09 ± 9.82) for symptomatic female and (33.2 ± 11.08) for asymptomatic ones. Significant high prevalence was detected in age group 28-37 years (45.5%). The overall prevalence of T. vaginalis in both was 27/200 (13.5%), in symptomatic (22%) and (5%) in asymptomatic ones. The main reported complaint of positive symptomatic cases was vaginal discharge (81.8%), lower abdominal pain and itching (63.6%), dyspareunia (50%) and dysuria (40.9%). No statistically significant difference between positive symptomatic and asymptomatic cases regarding age, marital status and residence (Table 1).
Table 1 Sociodemographics of positive symptomatic and asymptomatic cases.
| Symptomatic Positive T.vaginalis n=22 n(%) |
Asymptomatic positive T. vaginalis n=5 n(%) |
Test of significance P value |
|
|---|---|---|---|
| Age Mean ± SD (min-max) |
35.09±9.82 (20.0-58.0) |
33.2±11.08 (19.0-50.0) |
t=0.381 P=0.71 |
| Age groups | |||
| 18-27 | 4 (18.2) | 1 (20.0) | MC P=0.82 |
| 28-37 | 10 (45.5) | 3 (60.0) | |
| 38-47 | 5 (22.7) | 0 (0.0) | |
| 48-58 | 3(13.6) | 1 (20.0) | |
| Marital status | |||
| Married | 21 (95.5) | 5 (100.0) | FET P=1.0 |
| Divorced | 1 (4.5) | 0 (0.0) | |
| Residence | |||
| Urban | 9 (40.9) | 2 (40.0) | FET P=1.0 |
| Rural | 13 (59.1) | 3 (60.0) | |
Z=Mann whitney test, FET: Fischer exact test, χ2= Chis-square test, MC:Monte Carlo test, * P value significant <0.05, SD= standard deviation
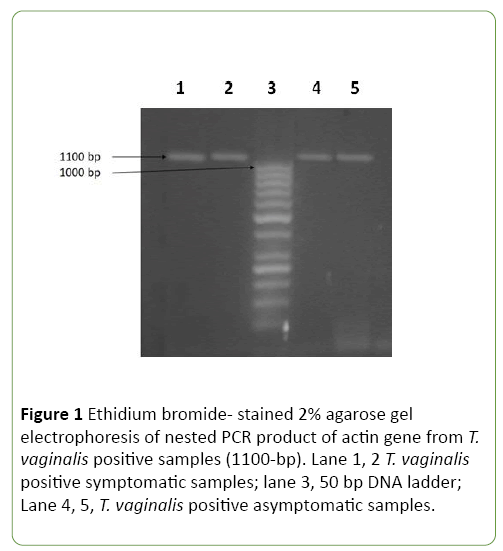
Thirteen positive T. vaginalis samples were detected (6.5%) by using wet mount method, while both culture and PCR methods detected 27 (13.5%) positive samples. All positive samples by culture were also positive by nested PCR and actin gene (1100 bp) was amplified from extracted DNA of all positive cases (Figure 1).
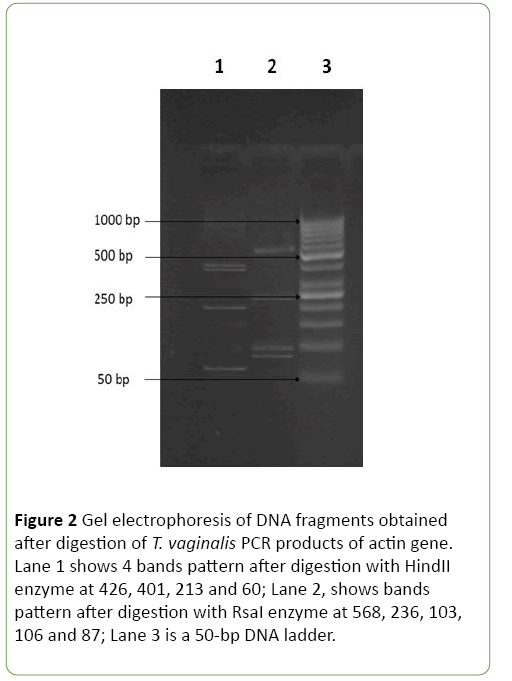
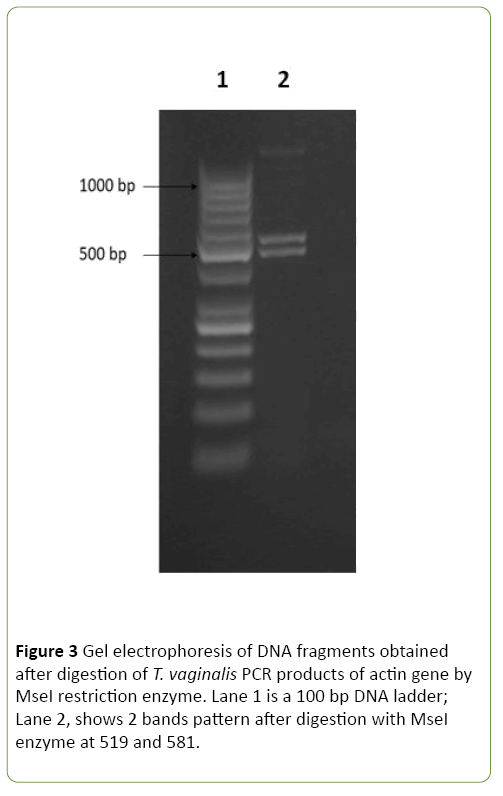
The amplified products of actin gene by nested PCR were digested by three restriction endonucleases including, HindII, MseI and RsaI. As shown in Figures 2 and 3 digestions of the amplified products with HindII restriction enzyme yielded four DNA bands at 426, 401, 213, and 60 bp. While, the restriction enzyme RsaI digested the PCR products into five bands at 568, 236, 106, 103, and 87 bp. Using MseI, two bands were detected at 519 and 581bp. On Crucitti et al. [15] as shown in Table 2, genotype H was defined as a representative of pattern 2 after HindII and MseI digestion and pattern 1 after RsaI digestion. Accordingly, actin genotype H is the only genotype detected in all positive 27 symptomatic and asymptomatic T. vaginalis isolates with no statistical difference between both groups.
Figure 2: Gel electrophoresis of DNA fragments obtained after digestion of T. vaginalis PCR products of actin gene. Lane 1 shows 4 bands pattern after digestion with HindII enzyme at 426, 401, 213 and 60; Lane 2, shows bands pattern after digestion with RsaI enzyme at 568, 236, 103, 106 and 87; Lane 3 is a 50-bp DNA ladder.
Table 2 Size of fragments, pattern groups and actin genotypes of the T. vaginalis[15].
| Genotype | Restriction with HindII (bp) | Restriction withRsaI (bp) | Restriction with MseI (bp) |
|---|---|---|---|
| A | 827,213, 60 | 568,236,190, 106 | 581, 519 |
| E | 827,213, 60 | 568,236,106,103, 87 | 581,315, 204 |
| G | 426, 401,213, 60 | 568,236,190, 106 | 581, 519 |
| H | 426,401,213, 60 | 568,236,106,103, 87 | 581, 519 |
| I | 426,401,213, 60 | 452,236,190,116, 106 | 581, 519 |
| M | 426,401,213, 60 | 568,236,190, 106 | 581,333, 186 |
| N | 426,401,213, 60 | 568,236,106,103, 87 | 581,333, 186 |
| P | 426,401,213, 60 | 452, 236, 116, 106, 103, 87 | 581,333, 186 |
Discussion
Trichomoniasis is highly prevalent, frequently asymptomatic, and simply communicable among sex partners [22,23]. Trichomoniasis is characterized by significant heterogeneity in its clinical presentation as pathogenicity, metronidazole sensitivity, sequence to infection, and susceptibility to acquisition of other infectious agents [24]. Whether this clinical variability is caused by host factors or to differences in the phenotypic expression of individual T. vaginalis isolates is not clear. However, recent advances in genetic characterization of T. vaginalis isolates showed that the extensive clinical variability in trichomoniasis and its disease consequences are matched by significant genetic diversity in the organism itself, suggesting a correlation between the genetic identity of isolates and their clinical manifestations [10]. A number of investigations using different genetic markers as well as different molecular techniques such as RFLP and microsatellite technique have been done [25]. But, this association is still not proven and so, we studied this association among females in our locality.
In the present study, the overall prevalence of T. vaginalis infection was 13.5%. Globally, prevalence estimates vary from 0.9%-80% [26,27]. In some countries, it had wide variation: 1.2% in Libya and Jordan [28], 3.2% in Turkey [29] and 28.1% in Saudi Arabia [30]. In Egypt, several studies were carried out and revealed variant prevalence among symptomatic women with T. vaginalis, 10.16% in Cairo [31], 11% in Benha [32] and 12.7% in Minia [33]. However, higher prevalence was recorded in Cairo (23%) [34], 38.37% in Mansoura [35] and 79.9% in Alexandria [36]. The discrepancy between our results and those detected by Hegazy MM et al. [35] may be due to large number of examined females and using other diagnostic methods.
Strain typing techniques are valuable methods to study the epidemiology and diversity of parasitic infections. PCR-RFLP technique based on the actin gene amplification was considered as a sensitive and discriminative method for strain typing of T. vaginalis clinical isolates. In previous studies, digestive patterns of the actin gene with each of restriction enzyme HindII, RsaI and MseI, detected 8 different genotypes as the major genotypes in Zambia and Kinshasa, with G and E as the most prevalent genotypes, respectively Crucitti T et al. [15]. Studying genetic diversity using PCR-RFLP technique targeting actin gene in our study, showed that genotype H is the only detected type in both symptomatic and asymptomatic groups with no difference between symptomatic or asymptomatic groups. This is in agreement with other studies which revealed no difference between symptomatic and asymptomatic groups using PCR-RFLP [37] and RAPD technique [9,13,26]. Also, in a study carried out by Rojas et al. [38], a genetic band of 490 bps was detected from T. vaginalis isolates collected from symptomatic cases of trichomoniasis and was absent in isolates from asymptomatic studied participant. On the other hand, genotyping of T. vaginalis, using PCR–RFLP by Crucitti et al. [15] and Momeni et al. [39], found eight different types (A, E, G, H, I, M, N, and P) and five types (G, E, H, I and mixed genotypes of G and E) respectively. The variation of results may be due to large sample size from wide regions with high prevalence rate.
Conclusion
In conclusion, PCR-RFLP technique is a dependable method to study different T. vaginalis genotypes in a population. Genotype H is the only genotype in the studied symptomatic and asymptomatic group concluding that no association is present between T. vaginalis genotypes and its clinical variability in our locality. Further studies on large scale are needed to prove this finding.
References
- Secor WE, Meites E, Starr MC, Workowski KA (2014) Neglected parasitic infections in the United States: trichomoniasis. Am J Trop Med Hyg 90:800-804.
- Poole DN, McClelland RS (2013) Global epidemiology of Trichomonasvaginalis. Sex Transm Infect 89:418–422.
- Byun JM, Jeong DH, Kim YN, Lee KB, Sung MS, et al. (2015) Experience of successful treatment of patients with metronidazole-resistant Trichomonasvaginalis with zinc sulfate: A case series. Taiwan J Obstet Gynecol 54:617-620.
- Mielczarek E, Blaszkowska J (2016)Trichomonasvaginalis: pathogenicity and potential role in human reproductive failure. Infection 44:447-458.
- Fraga J, Rojas L, Sariego I, Fernández-Calienes A (2011) Double-stranded RNA viral infection of Trichomonasvaginalis and correlation with genetic polymorphism of isolates. Exp Parasito l127:593-599.
- Patil MJ, Nagamoti JM, Metgud SC (2012) Diagnosis of Trichomonasvaginalis from vaginal specimens by wet mount microscopy, in pouch TV culture system, and PCR. J Glob Infect Dis 4:22.
- Donders GG, Depuydt CE, Bogers JP, Vereecken AJ (2013)Association of Trichomonasvaginalis and cytological abnormalities of the cervix in low risk women.PLoS One 8:e86266.
- Vanacova S, Tachezy J, Kulda J, Flegr J (1997) Characterization of trichomonad species and strains by PCR fingerprinting. J Euk Microbiol 44:545-552.
- Kaul P, Gupta I, Sehgal R, Malla N (2004)Trichomonasvaginalis: random amplified polymorphic DNA analysis of isolates from symptomatic and asymptomatic women in India. Parasitol Int 53:255-262.
- Meade JC, Carlton JM (2013) Genetic diversity in Trichomonasvaginalis. SexTransm Infect 89:444-448.
- HamplV, Vanácová S, Kulda J, Flegr J (2001) Concordance between genetic relatedness and phenotypic similarities of Trichomonasvaginalis strains. BMCEvol Bio l1:11.
- Conrad MD, Gorman AW, SchillingerJA, Fiori PL, Arroyo R, et al. (2012) Extensive genetic diversity, unique population structure and evidence of genetic exchange in the sexually transmitted parasite Trichomonasvaginalis. PLoS Negl Trop Dis 6:1573.
- Simoes-Barbosa A, Lobo TT, Xavier J, Carvalho SE, Leornadecz E (2005)Trichomonasvaginalis: intrastrain polymorphisms within the ribosomal intergenic spacer do not correlate with clinical presentation. Exper Parasit 110:108-113.
- Carlton JM, Hirt RP, Silva JC, Delcher AL, Schatz M, et al. (2007) Draft genome sequence of the sexually transmitted pathogen Trichomonasvaginalis. Science 315:207-212.
- Crucitti T, Abdellati S, Van Dyck E, Buve A (2008) Molecular typing of the actin gene of Trichomonasvaginalis isolates by PCR-restriction fragment length polymorphism. Clin Microbiol Infect 14:844-852.
- Drouin G, De Sa MM, Zuker M (1994) The Giardia lamblia actin gene and the phylogeny of eukaryotes. J Mol Evol 41:841-849.
- Meade JC, de Mestral J, Stiles JK, Secor WE, Finley RW, et al. (2009) Genetic diversity of Trichomonasvaginalis clinical isolates determined by EcoRI restriction fragment length polymorphism of heat-shock protein 70 genes. Am J Trop Med Hyg 80:245-251.
- Singh B (1997) Molecular methods for diagnosis and epidemiological studies of parasitic infections. Int J Parasito l27: 1135-1145.
- Bradic M, Warring SD, Tooley GE, Scheid P, Secor WE, et al. (2017) Genetic Indicators of Drug Resistance in the Highly Repetitive Genome of Trichomonasvaginalis. Gen Biol Evo l9:1658-1672.
- McCann JS (1974) Comparison of direct microscopy and culture in the diagnosis of trichomoniasis. Br J Vener Dis 50:450-452.
- Espinosa N, Hernández R, Lopez-Griego L, Arroyo R, Lopez-Villasenor I (2001) Differences between coding and non-coding regions in the Trichomonasvaginalis genome: an actin gene as a locus model. Actatropica 78:147-154.
- Dharma MN, Umashankar KM, Sudha, Abed GN, Kavitha G (2013) Prevalence of the Trichomonasvaginalis infection in a tertiary care hospital in rural Bangalore, Southern India. J Clin Diag Resear 7:1401-1403.
- Meites E, Gaydos CA, Hobbs MM, Kissinger P, Nyirjesy P, et al (2015)A review of evidence-based care of symptomatic trichomoniasis and asymptomatic Trichomonasvaginalis infections. Clin Infect Disea 61: S837-S848.
- PetrinD, Delgaty K, Bhatt R, Garber G (1998) Clinical and microbiological aspects of Trichomonasvaginalis. Clin Microbiol Rev 11:300-317.
- Conrad M, Zubacova Z, Dunn LA, Upcroft J, Sullivan SA, et al (2011) Microsatellite polymorphism in the sexually transmitted human pathogen Trichomonasvaginalis indicates a genetically diverse parasite. Molbiochem Parasitol 175:30-38.
- Valadkhani Z, Assmar M, Esfandiari B, Amirkhani A, Hassan N, et al. (2008)Trichomoniasis in asymptomatic patients. Iran J Pub Healt 37:113-117.
- Javanbakht M, Stirland A, Stahlman S, Smith LV, Chien M, et al. (2013) Prevalence and factors associated with Trichomonasvaginalis infection among high-risk women in Los Angeles. Sex transm dis 40:804.
- Kassem HH, Majoud O A (2006)Trichomoniasis among women with vaginal discharge in Benghazi city, Libya. J Egy Soci Parasitol 36: 1007-1016.
- Selvitopu A, Ozcelik S,Değerli S (2005) The incidence of Trichomonasvaginalis in vaginal specimens from gynecologic patients. Turk Soci Parasitol 30:175-177.
- Mahafzah AM, Al-Ramahi MQ, Asa'd AM, El-Khateeb MS (2008) Prevalence of sexually transmitted infections among sexually active Jordanian females. Sex Transm Dis 35:607-610.
- Zaki MA, Moussa HM, Hassanin OM (2001) Evaluation of the OSOM Trichomonas rapid test for detection of trichomoniasisvaginalis. Parasite Uni J 4:177-1784.
- Hussein AH, Saleh MH, Nagaty IM, Ghieth KA, El-Azab NA (2015) Prevalence, clinical criteria and sociodemographic predictors of Trichomonasvaginalis Infection in Suspected Egyptian Women, using direct diagnostic techniques. Iran J Parasitol 10: 432-440.
- GabrNS, Kamal AM, Mohamed RT, AbdelwahabSF (2006) Sensitivity and specificity of wet mount, culture and PCR in diagnosing T. vaginalis infection in females attending the gynecology clinic of Minia University Hospital. Minia Med Bull 17:1.
- Elsherif RH, Youssef MAF (2013) Real-time PCR improve detection of Trichomonasvaginalis compared to conventional techniques. Com Clin Pathol 22:295-300.
- Hegazy MM, El-Tantawy NL, Soliman MM, El-Sadeek ES, El-Nagar HS (2012) Performance of rapid immunochromatographic assay in the diagnosis of Trichomoniasisvaginalis. Diagn Microbiol Infect Dis 35: 49-53.
- Negm AY, El-Haleem DA (2004) Detection of trichomoniasis in vaginal specimens by both conventional and modern molecular tools. J Egypt So cParasitol 34:589-600.
- Sapru K, Mohan K, Gupta I, Ganguly NK, Mahajan RC, et al. (1994) DNA banding patterns of Trichomonasvaginalis strains isolated from symptomatic and asymptomatic subjects. J Protozool Res 4:40-47.
- Rojas L, Fraga J, Sariego I (2004) Genetic variability between Trichomonasvaginalis isolates and correlation with clinical presentation. Infection. Gene Evolu 1:53-58.
- Momeni Z, Sadraei J, Kazemi B, Dalimi A (2015) Molecular typing of the actin gene of Trichomonasvaginalis isolates by PCR-RFLP in Iran. Exper Parasitol 159:259-263.
Open Access Journals
- Aquaculture & Veterinary Science
- Chemistry & Chemical Sciences
- Clinical Sciences
- Engineering
- General Science
- Genetics & Molecular Biology
- Health Care & Nursing
- Immunology & Microbiology
- Materials Science
- Mathematics & Physics
- Medical Sciences
- Neurology & Psychiatry
- Oncology & Cancer Science
- Pharmaceutical Sciences