Md. Sajjad Hossain, Farzana Khanm Camellia, Nayon Uddin, Laila Arjuman Banu, Md. Kudrat-E-Zahan, Md. Masuqul Haque*
Department of Chemistry, University of Rajshahi, Rajshahi-6205, Bangladesh
Corresponding Author:
Md. Masuqul Haque
Department of Chemistry
University of Rajshahi
Rajshahi-6205, Bangladesh
E-mail: masuqul2003@ru.ac.bd
Received Date: March 05, 2019; Accepted Date: March 25, 2019; Published Date: April 02, 2019
Citation: Hossain MS, Banu LA, E-Zahan MK, Haque MM (2019) Synthesis, Characterization and Biological Activity Studies of Mixed Ligand Complexes with Schiff base and 2,2'-Bipyridine. Appl Sci Res Rev Vol.6 No.1:2 doi:10.21767/2394-9988.100086
Copyright: © 2019 Hossain MS, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Keywords
Schiff base ligands; Transition metals;2, 2'-Bipyridine; Isoniazid; Antibacterial activity
Introduction
Study of coordination chemistry of transition metal ions with various types of ligands has been enhanced by the current advancements in the fields of bioinorganic chemistry and medicine [1]. Through the years, Schiff bases have played a special role as chelating ligands in main group and transition metal coordination chemistry, due to their stability under a variety of oxidative and reductive conditions, and to the fact that imine ligands are borderline between hard and soft Lewis bases [2-4].
Schiff bases that arise from amino and carbonyl compounds are an important class of ligands that coordinate to metal ions through azomethine nitrogen and have been studied widely [5]. In azomethine derivatives, the (CN) linkage is necessary for biological activity; numerous azomethines have been reported to have marked antifungal, antibacterial, anticancer and antimalarial activities [6-14]. A large number of Schiff bases and their complexes have been studied for their interesting and significant properties, e.g. their ability to reversibly bind oxygen [15,16], catalytic activity in hydrogenation of olefins [17] photochromic properties [18] and complexing ability towards some toxic metals [19].
Mixed ligand complexes also play an important role in the biological field as exemplified by many ways in which enzymes are known to be activated by metal ions [20]. Schiff bases derived from heterocyclic compounds such as P- anisaldehyde and furan? 2?carbaldehyde have attracted an increased interest in the bioinorganic chemistry field [21-23]. Isoniazid (INH), also known as isonicotinylhydrazide (INH), is a known tuberculostatic agent used as a first-line agent for the prevention and treatment of both latent and active tuberculosis. It forms metal chelates with many bivalent ions having moderate to better biological significance [24,25]. Since the discovery of bipyridine at the end of the nineteenth century, the bipyridine ligand has been used extensively in the complexation of metal ions due to its strong redox stability and ease of functionalization [26,27]. The biological activities of the Schiff base and its mixed ligand complexes were also investigated herein.
Experimental
All used chemicals were purchased from Merck and Loba chemicals. All the melting points were determined on a digital melting point apparatus. Products were characterized by comparison of spectroscopic data (UV-Visible and FT-IR) and melting points with authentic samples. The wavelength of absorbance was determined by UV-Visible spectrophotometer [JASCO 503] using a quartz cuvette and ethanol as the reference. The IR spectrums were recorded on FT-IR spectrophotometer [JASCO, FT-IR/4100] Japan using dry KBr as the standard reference. The magnetic susceptibility of the complexes was measured at room temperature using a Gouy balance.
General Procedure for Synthesis of N-(4 methoxybenzylidene) isonicotinohydrazone Schiff base Ligand as the Primary Ligand, L1
INH (1.37 g, 10.0 mmol) was mixed with absolute ethanol (15 mL) and the mixture brought to the boil, producing a slurry. Barely sufficient additional ethanol was then added to give a homogeneous solution at reflux. P-anisaldehyde (1.22 mL, 10 mmol) was added drop-wise over 5 minutes and washed with 5 mL of ethanol. The reaction mixture was refluxed for 4 hours then allowed to cool slowly and to stand overnight. Finally, it produces a white crystalline solid which was filtered off and dried (Scheme 1).
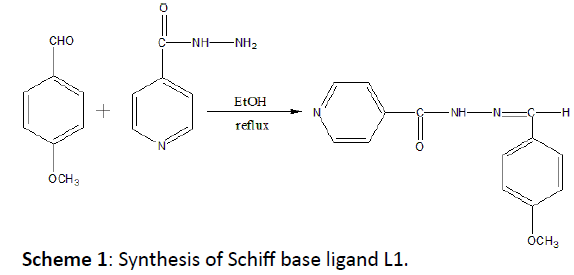
Scheme 1: Synthesis of Schiff base ligand L1.
Experimental procedure for synthesis of mixed ligand complexes
To the warm methanolic solution (10 mL) of primary ligand L1 (0.255 g, 1 mmol), 10 mL warm methanolic solution (0.257 g, 1 mmol) of nitrate salt of metal Cu(II) and Ni(II) were added. After 30 minutes 5 mL warm methanolic solution of 2,2′?bipyridine (0.156 g, 1 mmol) was added drop-wise as a secondary ligand (L2) and the resulting mixture was refluxed for about 3-4 hours. The obtained precipitate was filtered, washed with methanol and dried under vacuum on anhydrous CaCl2.
Antimicrobial activity
The ligand (L1) and its mixed ligand complexes with (L2) were screened for in vitro antimicrobial activity in DMSO against gramnegative Escherichia coli (E. coli) and gram-positive Bacillus cereus (B. cereus) strains by Kirby Bauer’s disc diffusion technique. A uniform suspension of test organism of 24 hours old culture was prepared in a test tube containing the sterile saline solution. Sterile nutrient agar was then added in each of the Petri dishes. The dishes were related to ensuring the uniform mixing of the microorganism in the agar medium which was then allowed to solidify. Sterile Whatmann filter paper discs were dipped in the solution of each compound and placed on the labeled plates. The DMSO was used as a control of the solvent. Kanamycin was used as a standard compound for comparison. Plates were kept in the refrigerator for half an hour for diffusion and incubated at 37°C for 24 hours. The diameter of the zone of inhibition around each disc was measured by scale and results were recorded in terms of mm. The observed data of antimicrobial activity of the L1, mixed ligand complexes and the standard drugs are given in Tables 4.
Results and Discussion
By the reaction of Cu(II) and Ni(II) with ligand (L1)and 2,2′? bipyridine (L2), complexes of the type [M(L1)(L2)]2+ were obtained. These complexes have a different color, stable at room temperature, insoluble in common polar solvent but soluble in DMSO and DMF, do not have the sharp melting point but decompose above 250°C. The measurement of molar conductivity at 10-3 M concentration carried out in DMSO at room temperature. The molar conductivity values show that the nitrate complexes were 1:2 electrolytes [28]. The analytical and physical data (color, melting point, molar conductivity and magnetic moment) of the complex are given in Table 1. For the Cu(II) and Ni(II) complexes the magnetic moments were 1.83 BM and 0.22 BM indicating paramagnetic and diamagnetic nature respectively. These values correspond to the square- planar geometry of both the mixed ligand complexes [29,30].
| Symbol of Compounds |
Complexes |
M.P or De (Decomposition Temp) / °C |
Solubility |
µeff in B.M |
| Color |
DMSO & DMF |
Molar conductance ohm-1 cm2mol-1 |
| Ligand (L1) |
C14H13N3O2 |
143 |
White |
(+) ve |
5 |
|
| [Cu(L1)(L2)] (NO3)2 |
[Cu(C14H13N3O2)(C10H8N2)] (NO3)2 |
275 (De) |
Brown |
(+) ve |
156 |
1.83 |
| [Ni(L1)(L2)] (NO3)2 |
[Ni(C14H13N3O2)(C10H8N2)] (NO3)2 |
285 (De) |
Yellow |
(+) ve |
149 |
Dia |
Table 1 Analytical and physical properties data of L1 and its mixed ligand complexes with L2.
IR spectral studies
The IR spectrum of the primary ligand (L1) showed in Table 2 and Figure 1, exhibited characteristic bands at 1658.84.5 cm-1 and 1598.69 cm-1 assigned to ?(C=O) and ?(C=N) respectively Figures 1-4 [31]. The band at 1658.84.5 cm-1 attributable to the ν(C=O) stretching vibration of the Schiff base ligand is shifted to another region 1628– 1637 cm-1 in the complexes of Cu(II) and Ni(II) indicating coordination of the carbonyl oxygen to the metal ions (Figures 2-3). The presence of band at 528– 535 cm-1 in the IR spectra of complex is due to M–O stretching vibration [28,32]. The azomethine band at 1598.69 cm-1 of Schiff base was shifted to lower frequency ranging of 1571–1593 cm-1 in the spectra of the complexes, confirming the participation of the azomethine nitrogen atom in the coordination of the metal ion. In the IR spectra of this complex, the new bands which appear in the 426– 434 cm-1 region are assigned to the ν(M–N) vibration [28,32]. The strong sharp band observed at 1384 cm-1 in the complex can be assigned to uncoordinated nitrate ion [28]. The band at 650 cm-1 is assigned to ν(C=N) of pyridine for secondary ligand, L2. This band is shifted to 671-682 cm-1 for mixed ligand complexes [33,34]. All of these IR data confirm ligands coordinated in Cu(II) and Ni(II) metal complexes through their O and N atoms respectively.
| Symbol of Compound |
Compound |
ν (C=O) |
ν (C=N) |
ρ(Py) bending |
V (M-O) |
ν (M- N) |
| Ligand (L1) |
C14H13N3O2 |
1658.84 |
1598.69 |
|
|
|
| Ligand L2 |
C10H8N2 |
|
1642 m |
650 |
|
|
| [Cu(L1)(L2)](NO3)2 |
[Cu(C14H13N3O2)(C10H8N2)] (NO3)2 |
1636.35 |
1592.95 |
678.73 |
528.02 |
433.99 |
| [Ni(L1)(L2)](NO3)2 |
[Ni(C14H13N3O2)(C10H8N2)] (NO3)2 |
1628.82 |
1571.41 |
681.68 |
534.71 |
426.92 |
Table 2 Key infrared bands (cm-1) of L1 and its mixed ligand complexes with L2.
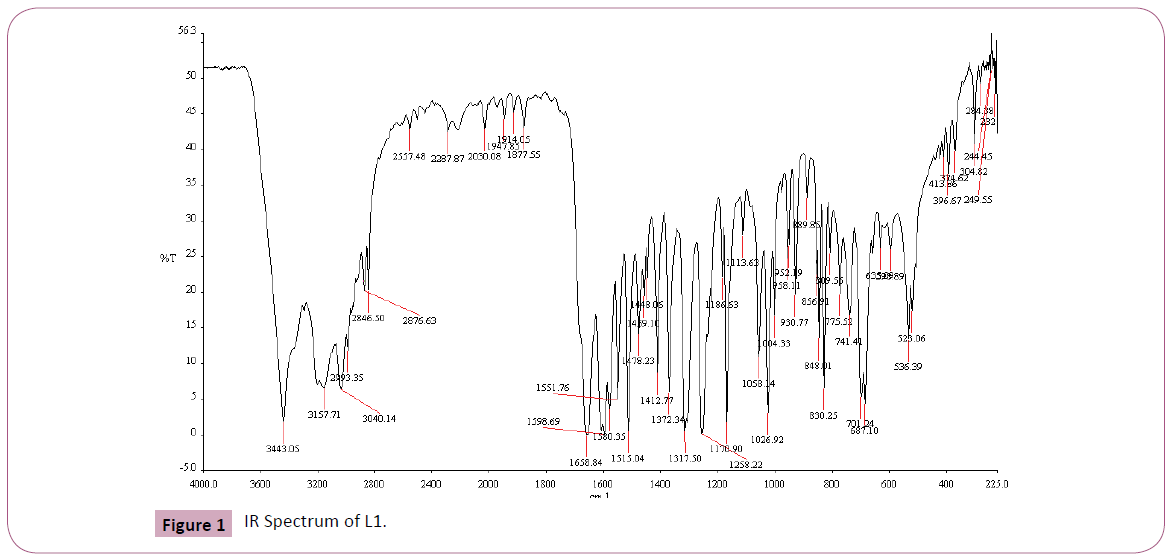
Figure 1: IR Spectrum of L1.
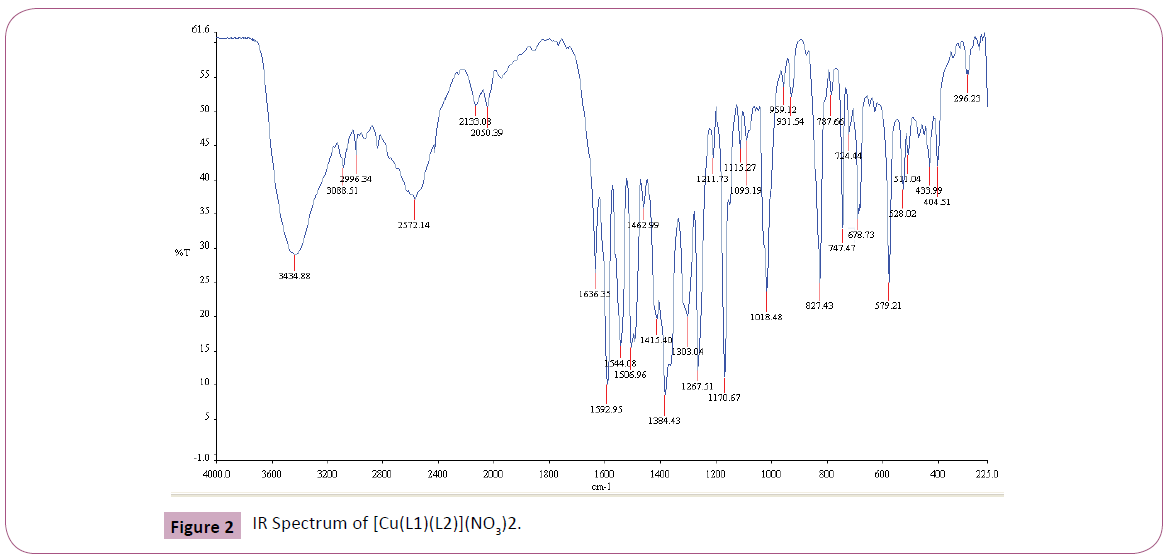
Figure 2: IR Spectrum of [Cu(L1)(L2)](NO3)2.
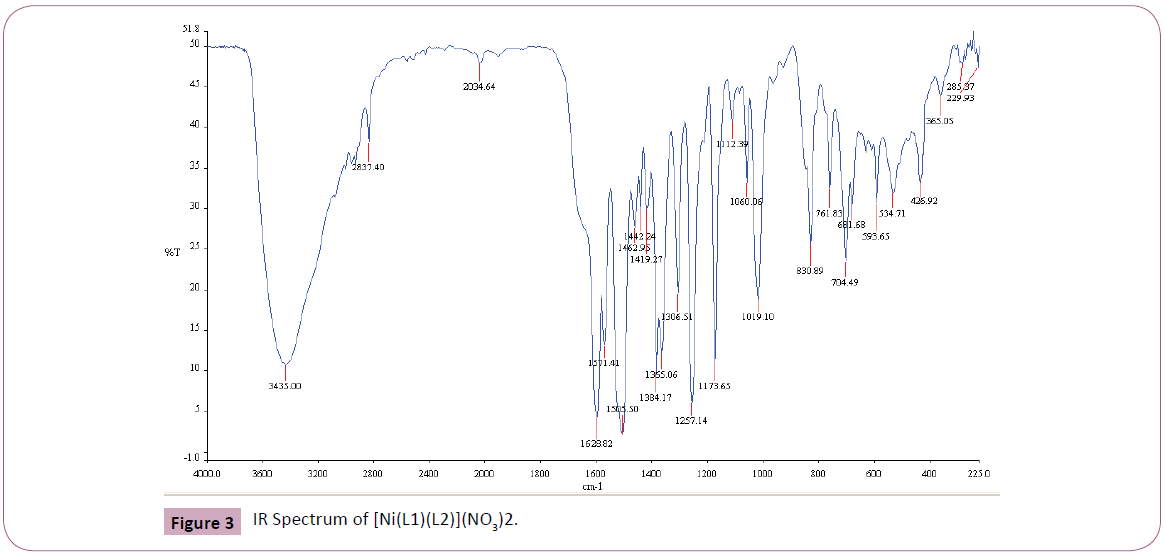
Figure 3: IR Spectrum of [Ni(L1)(L2)](NO3)2
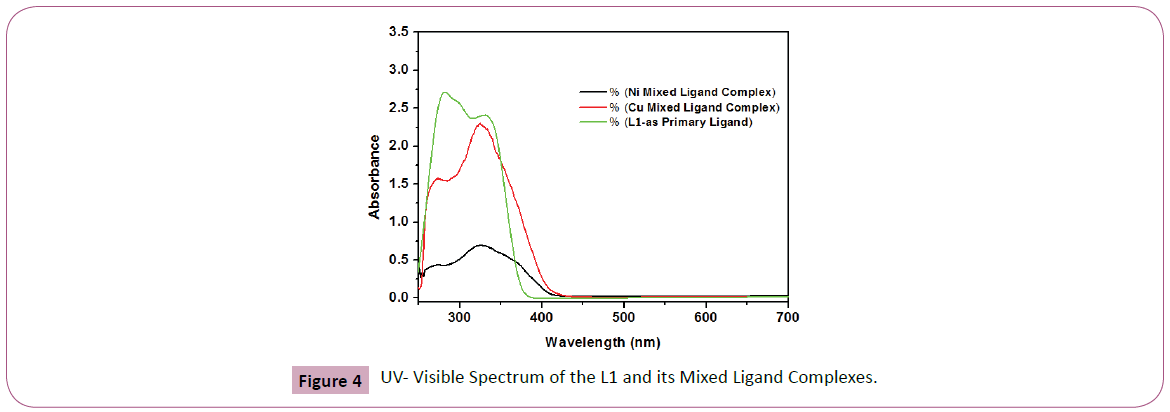
Figure 4: UV- Visible Spectrum of the L1 and its Mixed Ligand Complexes.
UV- visible spectra
The UV-Visible spectra of the ligand L1 shows two bands at 281 and 347 nm which are assigned to π- π* and n-π* transition respectively. All the complexes showed the charge transfer transitions which can be assigned to charge transfer from the ligand to metal (LMCT) and vice versa. For complexes, absorption bands at the range of 367– 370 nm may be associated with L→M charge transfer and vice versa (M→L ) [29,30]. In the UV-region, the complexes showed absorption band at 269–270 nm (Figure 4) which may be assigned to π −π * transition. The spectra of all the complexes exhibiting bands assigned to π −π and M→L charge transfer, the metals normally prefer squareplanar geometry [29,30]. All observations were summarized in the Table 3.
| Symbol of Compound |
Compound |
λ in nm |
Assignment |
| Ligand (L1) |
C14H13N3O2 |
283
333 |
π-π*
n-π* |
| [Cu(L1)(L2)](NO3)2 |
[Cu(C14H13N3O2)(C10H8N2)] (NO3)2 |
270
367 |
π -π*
C.T |
| [Ni(L1)(L2)](NO3)2 |
[Ni(C14H13N3O2)(C10H8N2)] (NO3)2 |
269
370 |
π -π*
C.T |
Table 3 UV- Visible spectrum of the Ligand L1 and its mixed ligand complexes with L2.
Antimicrobial screening result
The Schiff-base ligand L1 and its mixed ligand complexes with L2 reported here were evaluated for antibacterial activity against Escherichia coli and Bacillus cereus. The values of zone inhibition were measured in millimeter. The data of antibacterial activities of ligand L1 and its mixed ligand complexes with L2 are given in Table 4.
| Antibacterial Zone of Inhibition (in mm) |
| Compounds |
Gram Negative |
Gram Positive |
| Escherichia coli |
Bacillus cereus |
| Kanamycin |
32 |
35 |
| Ligand (L1) |
4 |
4 |
| [Cu(L1)(L2)](NO3)2 |
20 |
22 |
| [Ni(L1)(L2)](NO3)2 |
18 |
15 |
Table 4 Antibacterial screening results of Ligand L1 and its mixed ligand complexes with L2.
Based on the above results the expected structures of the Schiff base complexes may be represented as shown in the Figure 5.
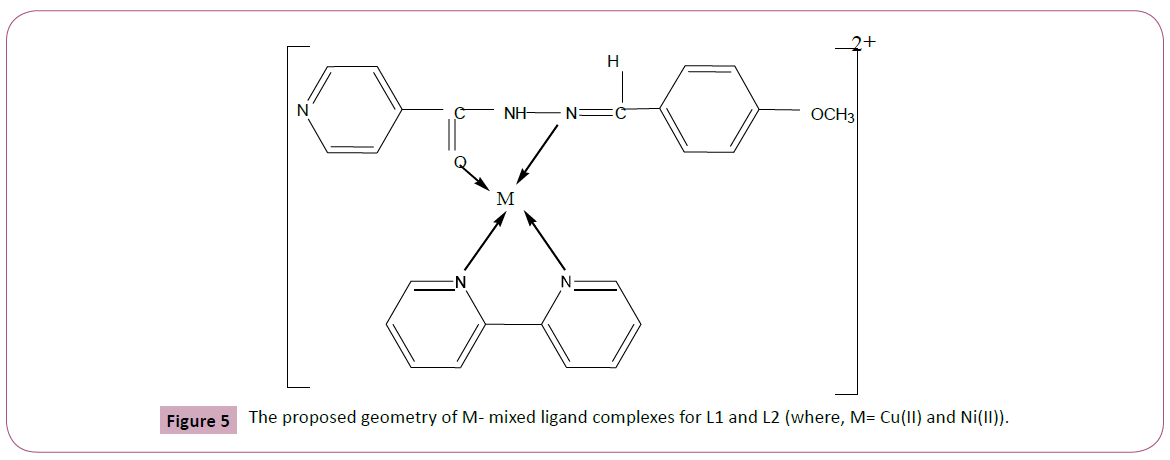
Figure 5: The proposed geometry of M- mixed ligand complexes for L1 and L2 (where, M= Cu(II) and Ni(II)).
The inhibitory zone data reveals that the ligand L1, as well as its mixed ligand complexes with L2, shows good to moderate good antibacterial activity. The biological activity of Schiff base ligand arises from the presence of imine group which imports in elucidating the mechanism of transformation reaction in biological systems. However, mixed ligand complexes with L2 showed remarkable antibacterial activity as a result of chelation of metal with organic ligands synergistically increasing its effect [28,31,32,35]. The DMSO control did not show any antimicrobial activity against the tested bacterial strains whereas the tested compounds were found to be active.
Conclusion
The Schiff base ligand L1 and its Cu(II) and Ni(II) mixed ligand complexes with L2 were prepared and characterized using various spectral and other techniques. The IR spectral data indicated that Schiff base (L1) and 2,2′?bipyridine (L2) ligand coordinated to the metal ions bycarbonyl oxygen, N of azomethine and pyridine respectively indicating the square- planar geometry of the complexes. It has been observed that the mixed ligand complexes showed more antibacterial activity than Schiff base ligand L1. Although with respect to standard, the tested compounds were found to be moderately active.
Acknowledgement
We are grateful thank to Department of Chemistry, Rajshahi University , Rajshahi, Bangladesh, for providing necessary materials and we are also grateful thank to Department of Pharmacy, Rajshahi University, Rajshahi, for providing Antibacterial activities.
References
- Hosseini SA, Samadzadeh P, Mirzaahmadi AA, Khandar AA, Mahmoudi G, et al. (2010) Synthesis, crystal structures, spectroscopic and electrochemical studies on Cu(II) and Ni(II) complexes with compartmental nitrogen–oxygen mixed donor ligands. Polyhedron 80: 41.
- Anacona J, Santaella J (2013) Synthesis, magnetic and spectroscopic studies of a Schiff base derived from cephaclor and 1,2-diaminobenzene and its transition metal complexes. Spectrochim Acta A 115: 800.
- Garnovskii AD, Nivorozhkin AL, Minkin VI (1993) Ligand environment and the structure of schiff base adducts and tetracoordinated metal-chelates. Coord Chem Rev 126: 1.
- Ziessel R (2001) Schiff-based bipyridine ligands. Unusual coordination features and mesomorphic behavior. Coord Chem Rev 216: 195.
- Yousif E, Majeed A, Sammarrae KA, Salih N, Salimon J, et al. (2013) Metal complexes of Schiff base: Preparation, characterization and antibacterial activity. Arab J Chem.
- Annapoorani S, Krishnan CN (2013) Synthesis and spectroscopic studies of trinuclear N4 schiff base complexes. Int J Chem Technol Res5: 180.
- Devidas SM, Quadri SH, Kamble SA, Syed FM, Vyavhare DY (2011) Novel one-pot synthesis of schiff base compounds derived from different diamine & aromatic aldehyde catalyzed by P 2O 5/SiO2 under free-solvent condition at room temperature. J Chem Pharm Res 3: 489.
- Xavier AL, Raj MA, Marie JM (2012) Synthesis and spectral characterization of an aminoacetophenone-based Schiff base and its interaction studies with ascorbic acid. J Chem Pharm Res 4: 669.
- Padmaja M, Pragathi J, Kumari CG (2011) Synthesis, spectral characterization, molecular modeling and biological activity of first row transition metal complexes with Schiff base ligand derived from chromone-3-carbaldehyde and o-amino benzoic acid. J Chem Pharm Res 3: 602.
- Thorat BR, Jadhav R, Mustapha M, Lele S, Khandekar D, et al. (2011) Synthesis and Fluorescence properties of Schiff bases of 4-hydroxy-3- methoxy-5-nitrobenzaldehyde. J Chem Pharm Res 3: 1045.
- Raman N, Pitchaikani RY, Kulandaisamy A (2001) Synthesis and characterization of Cu(II), Ni(II), Mn(II), Zn(II) and VO(II) Schiff base complexes derived fromo-phenylenediamine and acetoacetanilide. J Chem Sci 113: 183.
- Mishra AP, Rajendra KJ (2010) Microwave synthesis, spectroscopic, thermal and biological significance of some transition metal complexes containing heterocyclic ligands. J Chem Pharm Res 2: 51.
- Freeman HC (1973) Metal Complex3s of amino acids and peptides. Inorga Biochem 1: 121.
- Ajay RP, Kamini JD, Sambhaji SR, Vishwanath RP, Rama SL (2012) Synthesis characterization and biological activity of mixed ligand Co(II) complexes of Schiff base 2-amino-4-nitrophenol-n-salicylidene with some amino acids. J Chem Pharm Res 4: 1413.
- Mohamed GG, Omar MM, Hindy AM (2006) Metal Complexes of Schiff Bases: Preparation, Characterization, and Biological Activity. Turk J Chem 30: 361.
- Jones RD, Summerville DA, Basolo F (1979) Synthetic oxygen carriers related to biological systems. Chem Rev 79: 139.
- Olie GH, Olive S (1984) The Chemistry of the Catalyzes Hydrogenation of Carbon Monoxide. Springer 152: 301.
- Margerum JD, Miller LJ (1971) Photochromic processes by Tautomerism in Photochromism. Wiley Intersci 569.
- Sawodny WJ, Riederer M (1977) Addition Compounds with Polymeric Chromium(II)?Schiff Base Complexes. Angew Chem Int Ed Eng 16: 859.
- Mahmoud WH, Mahmoud NF, Mohamed GG, El?Sonbati AZ, El?Bindary AA (2015) Synthesis, spectroscopic, thermogravimetric and antimicrobial studies of mixed ligands complexes. J Mol Struct 1095: 15.
- Chaviara AT, Cox PJ, Repana KH, Papi RM, Papazisis KT, et al. (2004) Copper(II) Schiff base coordination compounds of dien with heterocyclic aldehydes and 2-amino-5-methyl-thiazole: synthesis, characterization, antiproliferative and antibacterial studies. Crystal structure of CudienOOCl2. J Inorg Biochem 98: 1271.
- Ciller JA, Seoane C, Soto JL, Yruretagoyena B (1986) Synthesis of heterocyclic compounds. LI†. Schiff bases of ethyl 2?aminofurancarboxylates. J Heterocycl Chem 23: 1583.
- Ali OAM (2014) Synthesis, spectroscopic, fluorescence properties and biological evaluation of novel Pd(II) and Cd(II) complexes of NOON tetradentate Schiff bases. Spectrochim Acta 121: 188.
- Zommer S, Lipiec T (1963) Determination of isonicotinic acid hydrazide in various substances and tablets with cu2+ ions in the presence of acetone. Acta Poloniae Pharmaceutica 20: 229.
- Munson JW, Connors KA (1972) Spectrophotometric determination of acid hydrazidesvianickel (II)? catalyzed hydroxamic acid formation. J Pharma Sci 61: 211.
- Kaes C, Katz A, Hosseini MW (2000) Bipyridine: the most widely used ligand. A review of molecules comprising at least two 2,2'-bipyridine units. Chem Rev 100: 3553.
- Blue F (1889) About the dry distillation of pyridinecarboxylic salts. Monatsh Chem 10: 375.
- Mariana LD, Angela K, Nicoale S, Adin MM (2010) Transition metal M(II) complexes with isonicotinic acid 2-(9-anthrylmethylene)-hydrazide. J Serb Chem Soc 75: 1515.
- Ikechukwu PE, Ajibade (2015) Synthesis, Characterization and Biological Studies of Metal(II) Complexes of (3E)-3-[(2-{(E)-[1-(2,4-Dihydroxyphenyl) ethylidene]amino}ethyl)imino]-1-phenylbutan-1-one Schiff Base. Mole 20: 9788.
- Raman N, Dhavethu RJ, Sakthivel AJ (2007) Synthesis, spectral characterization of Schiff base transition metal complexes: DNA cleavage and antimicrobial activity studies. Chem Sci 119: 303.
- Ganesh SK, Priyanka LA, Digambar DG, Dnyandeo MJ (2016) Efficient ultrasound synthesis and biology screening of metal complexes of N-(4- methoxybenzylidene) Isonicitinohydrazone. W J Pharma Research 5: 1153.
- Abdul H, Ahmed A, Salima A, Ben G (2008) Synthesis And Characterization of Some Transition Metals Complexes of Schif Base Derived From Benzidine and Acetylacetone. J Science and its Appl 2: 83.
- Mohamed GG, Omar MM, Ibrahim AA (2009) Biological activity studies on metal complexes of novel tridentate Schiff base ligand. Spectroscopic and thermal characterization. Eur J Med Chem 44: 4801.
- Abd EH, Omar MM, Mohamed GG (2011) Synthesis, structural, thermal studies and biological activity of a tridentate Schiff base ligand and their transition metal complexes. Spectrochim Acta 78: 36.
- Vogt RL, Dippoid L (2005) Escherichia Coli O157: H7 Outbreak Associated with Consumption of Ground Beef, June–July 2002. Pub Health Rep 120: 174.