Babak Abdolkarimi1*, Javad Dehbozorgian2 and Shahrzad Razmjooee2
1Department of Pediatrics, Lorestan University of Medical Sciences, Iran
2Department of Pediatrics, Shiraz University of Medical Sciences, Iran
Corresponding Author:
Abdolkarimi B
Assistant Professor of Pediatric Hematology and Oncology Department
Lorestan University of Medical Sciences, Iran
Tel: +9183605274
E-mail: b.abdolkarimi@yahoo.com
Received Date: September 26, 2017; Accepted Date: October 18, 2017; Published Date: October 26, 2017
Citation: Abdolkarimi B, Dehbozorgian J, Razmjooee S (2017) Scalar Scoring in Thalassemia Genotype: As a New Overview. Appl Sci Res Rev 4:9. doi: 10.21767/2394-9988.100060
Introduction
One of the most complex subjects of Hematology is thalassemia and hemoglobinopathy genotype interpretation and phenotype determination as well as their clinical symptoms and electrophoresis. Therefore, establishing a logical digital relationship between the above-mentioned items can contribute to the understating of this subject.
In this opinion article, we present a new digital scoring method for genotype and predicting phenotype and some CBC indexes based on estimated genotype score. This method in Hematology might facilitate the complexity of medical patients, but it may not be possible to justify all existing details in these patients.
Material and Methods
Focusing on down table: For example, a 6-year-old patient with βA/ β+ genotype is a minor β thalassemia case and he has β score=4 and α score=6 and total score=10 in our scoring. In this method, we attribute his minimum Hb as a sum of α score and β score equal 10 and his minimum MCV=76. Also, in this patient because of β thalassemia minor, genotype is asymptomatic with predictable Hb electrophoresis.
The thalassemia has Mendelian inheritance and it has inherited autosomal recessive; therefore, people with 50% healthy alleles have thalassemia trait and they are not patients, but the people with 100% involved alleles of α or β genes have thalassemia disease and they are patients. Thalassemic patients have organomegaly, anemia, and ineffective erythropoiesis with objective and subjective symptoms (fatigue, pallor, jaundice) and abnormal Hb electrophoresis and CBC, but thalassemia trait persons just have abnormal Hb electrophoresis and CBC without clinical symptoms (Figure 1).
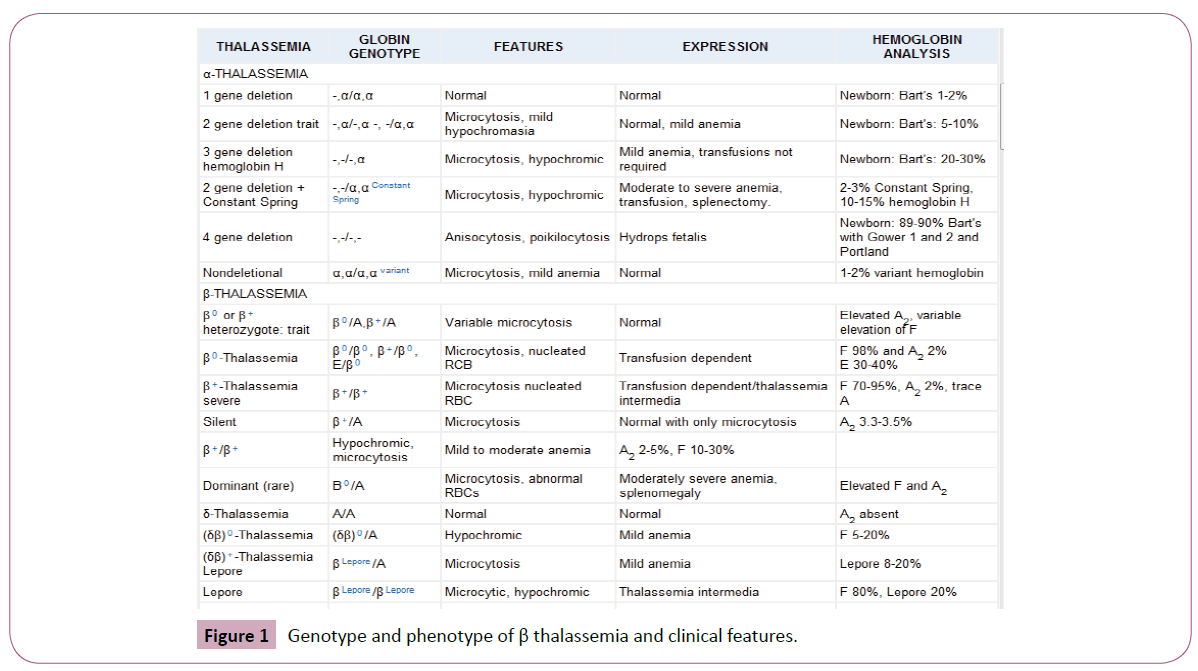
Figure 1: Genotype and phenotype of β thalassemia and clinical features.
In this method, we attributed β chain score equal 3.
βA, β+, β0 and 6 compound genotypes derive from this three conditions. Every healthy β chain gets the score +3: HbA has α2β2 combination; therefore, a healthy person with two healthy β chains in hemoglobin A is shown as βA/βA has β score=+6, So: (Figure 2a-c).
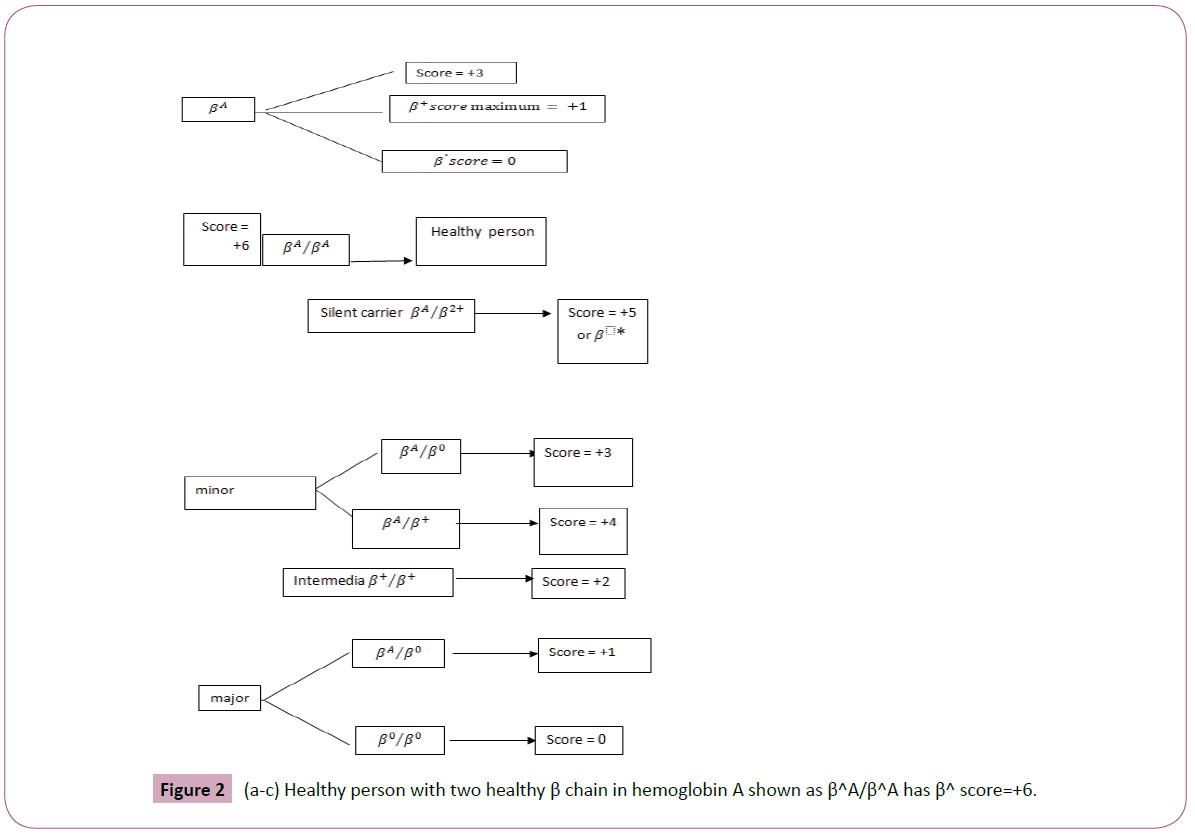
Figure 2: (a-c) Healthy person with two healthy β chain in hemoglobin A shown as β^A/β^A has β^ score=+6.
Hemoglobin Lepore is a chimeric protein, which is resulted from a fusion gene (adherence of β gene to Δ). Therefore: (Figure 3).
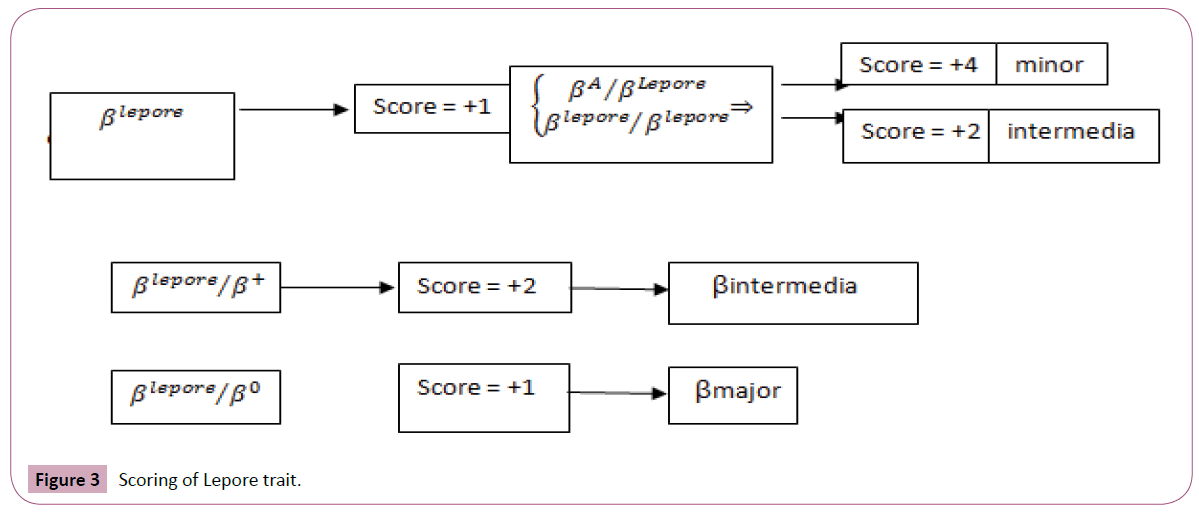
Figure 3: Scoring of Lepore trait.
Based on the above description, Lepore trait has β score=+4; therefore, the patient has minor thalassemia and Lepore homozygous patient has β score=+2 and the patient has intermediate thalassemia. With the same method composition of βlepore with other β gene variations such as βE, βC, βS, βD produced other β thalassemia and hemoglobinopathies genotypes and phenotypes.
Exception 1: βlepore/ βlepore (homozygote Lepore or Lepore disease) is one of exceptions, which has maximum 20-25% Lepore Hb with score=2. Therefore, he is an intermediate β thalassemia phenotype or disease. This patient has 75-80% HbF. People with Lepore heterozygote genotype such as βlepore βA have score=4; therefore, they are β variant thalassemia trait and have no disease. In these cases, HbA1 and HbA2 do not exist and remaining Hb chains are 75-80% δ chains that present with HbF.
Note: Second interesting genotype in β thalassemia is co-deletion of β gene and Δ. One of this genotypes is (Δβ)+/ (Δβ)+. This person has β score=+2 and he is β thalassemia intermediate with low HbA2, HbA1. (Δβ)+/A is heterozygous with score=+4; therefore, he is a case of β thalassemia trait or minor β thalassemia with low HbA2.
Genotype and phenotype multiplication table: In summary, if we put all β genes situations in horizontal and vertical column, we assess all probable β genotypes. Figure 1 shows possible genotypes with β gene variations, Figure 3 shows possible genotypes score in our scalar scoring, and Figure 4 shows possible clinical situations compatible with genotype and score. For example, patient with β+/ β+ in matrix home 4 x 4 has β thalassemia intermediate genotype and β score=+2 in our scoring (Figures 4-6).
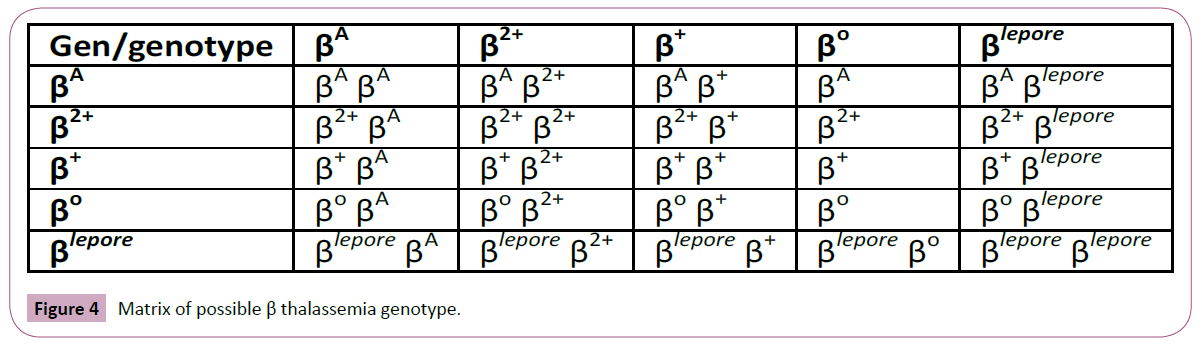
Figure 4: Matrix of possible β thalassemia genotype.
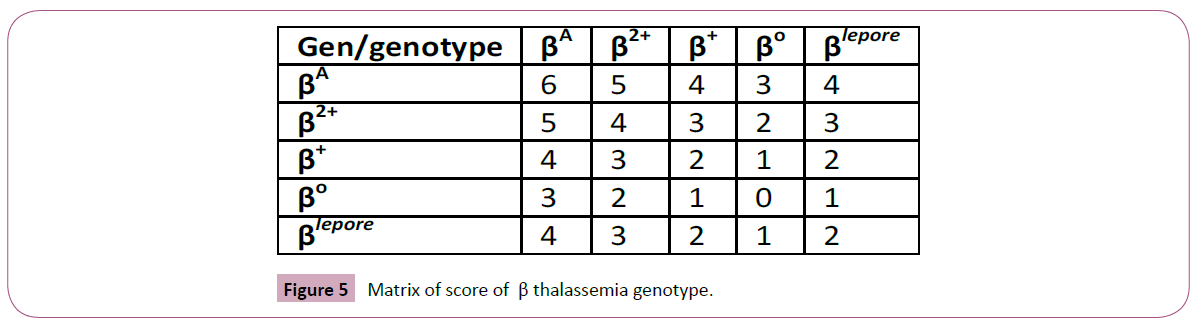
Figure 5: Matrix of score of β thalassemia genotype.
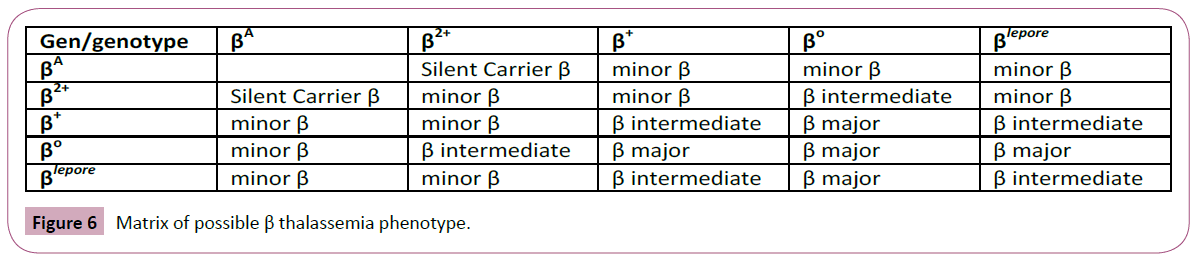
Figure 6: Matrix of possible β thalassemia phenotype.
α genes placement and the score in αthalassemia: There are two types of α genes: α1 and α2 genes and they are not equal regarding their ability for producing α chains. Therefore, they have 2 different scalar scores in our scoring system. In healthy persons, α genes composition is α2 α1\α2 α1. If we attribute α1 score=+1 and α2 score=+2 in scoring, they have 2 different genotypes with α genes in α thalassemia setting: (Figure 7).
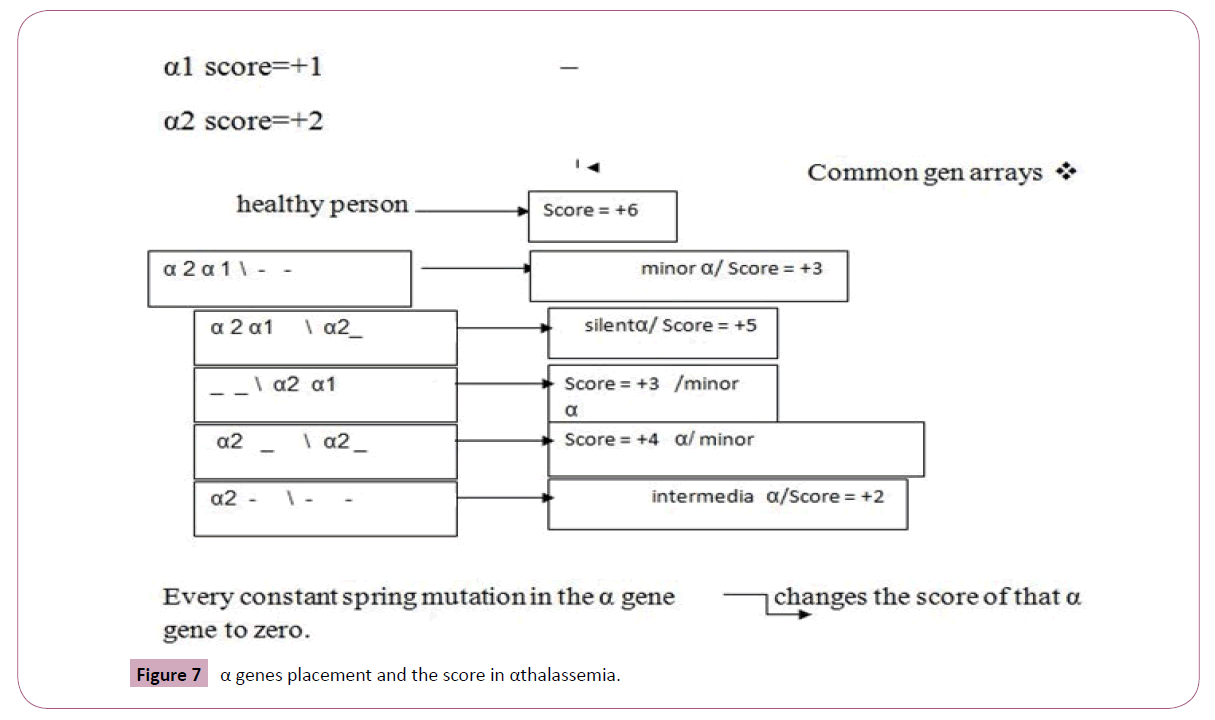
Figure 7: α genes placement and the score in αthalassemia.
This method is extended about α variants hemoglobinopathies such as Constant Spring Hemoglobin and other deletional and non-deletional genotypes:
α CS score has value 0 in our scoring system, for example: α2 _ \ α CS_ has score=+2, therefore, he is intermediate α thalassemia.
Estimation of complex genotype clinical manifestations: In complex genotype such as α thalassemia intermediate with β thalassemia minor scor α=+2 and β thalassemia minor score=+3 or +4 and α/β ratio=2/3 or 4. In pure genotype such as α thalassemia intermediate with β normal scor α=+2 and β score=+6 and α/β ratio=2/6. In comparison of two patients, α/β ratio nearer to 1 (normal genotype score in healthy person) clinical manifestations would be better.
Conclusion
Our new and simplified method in interpreting thalassemia genotype, phenotype, and clinical symptoms well based on the scoring method. Therefore, it is an effective applied method for learning, interpreting, and simplifying complicated concepts of thalassemia syndromes. We think that using simple language is most important factor for effectiveness of this course and this method similar case based bedside teaching provides active learning in real context observes students skills, increases learners' motivation and professional thinking, integrates clinical, communication, problem solving, decision making and ethical skills [1-5].
References
- Kliegman R, Stanton B, St. Geme J, Schor N, Behrman RE (2011) 19th edition, Nelson Textbook of Pediatrics. Elsevier Saunders, Philadelphia, USA.
- Al-Allawi NAS, Jalal SD, Mohammad AM, Omer SQ, Markous RSD (2014) β-Thalassemia Intermedia in Northern Iraq. Hindawi Publishing Corporation, Egypt.
- Rahim F (2008) Genotyping of Thalassemia in Microcytic Hypochromic Anemia Patients from Southwest Region of Iran. Pak J Med Sci 24: 23-28.
- George E, Ann TJ (2011) Genotype-Phenotype Diversity of Beta-Thalassemia in Malaysia: Treatment Options and Emerging Therapies. Med J Malaysia 65: 256-260.
- Shahriari M (2014) Case Based Teaching at the Bed Side Versus in Classroom for Undergraduates and Residents of Pediatrics. J Adv Med Educ Prof 2: 135-136.