Tafe LJ, de Abreu FB, Peterson JD and Tsongalis GJ*
Laboratory for Clinical Genomics and Advanced Technology (CGAT), Department of Pathology and Laboratory Medicine, Dartmouth- Hitchcock Medical Center and the Norris Cotton Cancer Center, Lebanon, NH, USA
*Corresponding Author:
Gregory J Tsongalis
Department of Pathology and Laboratory
Medicine, Dartmouth-Hitchcock Medical
Center, 1 Medical Center Drive, Lebanon
NH, USA
Tel: +16036505498
Fax: +16036507214
E-mail: Gregory.j.tsongalis@hitchcock.org
Received date: December 07, 2016; Accepted date: December 12, 2016; Published date: December 15, 2016
Citation: Tafe LJ, de Abreu FB, Peterson JD, et al. MET Exon 14 Skipping Mutation in Non-Small Cell Lung Cancer Identified by Anchored Multiplex PCR and Next-Generation Sequencing. J Cancer Epidemiol Prev. 2016, 1:1.
Recurrent MET exon 14 (MET ex14) somatic splice site mutations have been described in 0.6 to 7% of lung non-small cell carcinomas (NSCLC) [1-3]. These mutations result in exon 14 skipping and subsequent MET activation with clinical trials demonstrating promising sensitivity to c-MET inhibitors [3]. These mutations are typically mutually exclusive of other lung adenocarcinoma known oncogenic driver mutations (e.g., EGFR, KRAS, BRAF, ERBB2, ALK, ROS1) but frequently co-occur with MDM2 and CDK4 amplification on chromosome 12q [2]. MET ex 14 mutations display diversity with upwards of 126 distinct DNA sequence variants described necessitating comprehensive genomic profiling by clinical laboratories for routine detection of these mutations in patients [2]. Of note, these MET ex14 splice site mutations are not limited to just NSCLC but also have been identified in a small percentage of gliomas and gastro-esophageal carcinomas [2,4].
Anchored multiplex PCR and next-generation sequencing is a novel method adopted by some clinical laboratories to detect somatic fusion events in formalin-fixed paraffin-embedded (FFPE) tumor tissues. Using RNA and DNA based NGS methods, we identified and confirmed a MET ex14 splice site mutation in two tumor tissues from an 82-year-old woman with lung adenocarcinoma. In our laboratory, DNA and RNA is extracted simultaneously from using the Qiagen AllPrep DNA/RNA FFPE Kit on the QIAcube. Next generation sequencing (NGS) is performed on both the RNA and DNA extracted from the FFPE. RNA undergoes reverse transcription to cDNA and is tested for fusions and other mutations associated with 53 genes using the Archer® FusionPlex® Solid Tumor Kit on the Illumina MiSeq instrument. One of the main advantages of the Archer assay’s chemistry (anchored multiplex PCR) is that it can detect fusions associated with the genes in this kit without prior knowledge of fusion partners or breakpoints by using universal adaptors at one end of the cDNA [5]. Following sequencing, data analysis is performed using Archer Analysis software (v4.0.11); at this point any fusion partner genes or sequence variants can be identified after aligning to the reference genome. DNA is also tested for hotspot mutations in 50 genes using the AmpliSeq v2 kit on the Ion Torrent PGM instrument; analysis is performed using Torrent Suite (v4.0.2) for alignment and variant calling and Golden Helix SVS (v8.3.4) for annotation. Testing our patient’s tumor DNA with the AmpliSeq v2 assay identified two genomic alterations, a TP53 mutation (c.527G>A, p.C176Y) in exon 5 and a MET substitution mutation (c.2942-1G>A) in the splice site acceptor region (3’ region of intron upstream of ex 14; genomic position 116,411,902) of exon 14 (Figure 1a). The MET ex14 mutation was confirmed with the Archer FusionPlex assay and identified as an oncogenic isoform of MET resulting in skipping of exon 14 (Figure 1b). This particular MET mutation has been previously described by Frampton et al. [2].
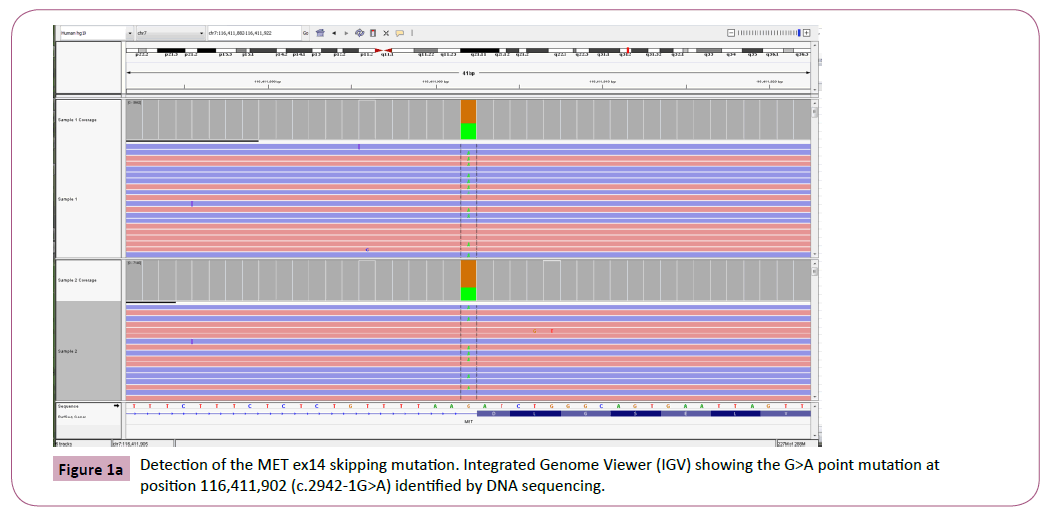
Figure 1a: Detection of the MET ex14 skipping mutation. Integrated Genome Viewer (IGV) showing the G>A point mutation at position 116,411,902 (c.2942-1G>A) identified by DNA sequencing.
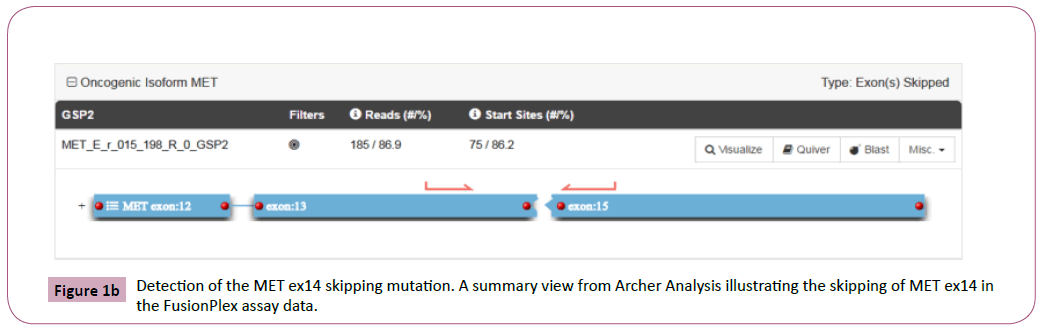
Figure 1b: Detection of the MET ex14 skipping mutation. A summary view from Archer Analysis illustrating the skipping of MET ex14 in the FusionPlex assay data.
Patients with MET ex14 skipping mutations are eligible for therapy with c-MET inhibitors and it is important for clinical laboratories to identify this population of patients. Anchored Multiplex PCR offers the ability to detect structural variants in multiple clinically significant genes, including MET ex14 mutations, and can be incorporated by clinical laboratories into the workflow for genotyping tumors. Several clinical trials are currently ongoing for patients with MET ex14 skipping mutations including: a phase II study in stage IIIB/IV lung adenocarcinoma with tepotinib who failed standard of care treatment (NCT02864992); a phase II study of capmatinib in NSCLC patients who have received prior MET inhibitor therapy (NCT02750215); NCI-MATCH trial has a phase II arm with crizotinib therapy (NCT02465060); a Phase II, i study of savolitinib (HMPL-504) for patients with locally advanced/ metastatic MET-mutation-positive pulmonary sarcomatoid carcinomas (NCT02897479); and, a phase I study of Pf-02341066, a c-met/hgfr selective tyrosine kinase inhibitor in patients with advanced cancer (PROFILE 1001) (NCT00585195). Response of MET ex14 skipping mutation positive tumors to c-MET inhibitors and eligibility for several clinical trials makes this exon skipping mutation a clinically actionable mutation with relevance to therapeutic strategy and outcomes.
In this study, we describe the detection of a MET 14 exon skipping mutation in two tissues from the same patient using both RNA and DNA sequencing. The importance of this approach is the absolute confirmation of this exon skipping mutation by first DNA sequencing and then RNA sequencing. Finally, accurate detection and confirmation of this mutation is required as it does have therapeutic implications for the patient.
Acknowledgements
The data presented in this manuscript was supported and generated through CGAT in the Department of Pathology and Laboratory Medicine of the Geisel School of Medicine at Dartmouth, the Dartmouth Hitchcock Medical Center, and the Norris Cotton Cancer Center.
References
- Schrock AB, Frampton GM, Suh J, Chalmers ZR, Rosenzweig M, et al. (2016) Characterization of 298 Patients with Lung Cancer Harboring MET Exon 14 Skipping Alterations. J Thorac Oncol 11: 1493-1502.
- Frampton GM, Ali SM, Rosenzweig M, Chmielecki J, Lu X, et al. (2015) Activation of MET via diverse exon 14 splicing alterations occurs in multiple tumor types and confers clinical sensitivity to MET inhibitors. Cancer Discov 5:850-859.
- Paik PK, Drilon A, Fan PD, Yu H, Rekhtman N, et al. (2015) Response to MET inhibitors in patients with stage IV lung adenocarcinomas harboring MET mutations causing exon 14 skipping. Cancer Discov5:842-849.
- Van Der Steen N, Giovannetti E, Pauwels P, Peters GJ, Hong DS, et al. (2016) cMET Exon 14 Skipping: From the Structure to the Clinic. J Thorac Oncol11:1423-1432.
- Zheng Z, Liebers M, Zhelyazkova B, Cao Y, Panditi D, et al. (2014) Anchored multiplex PCR for targeted next-generation sequencing. Nat Med20:1479-1484.